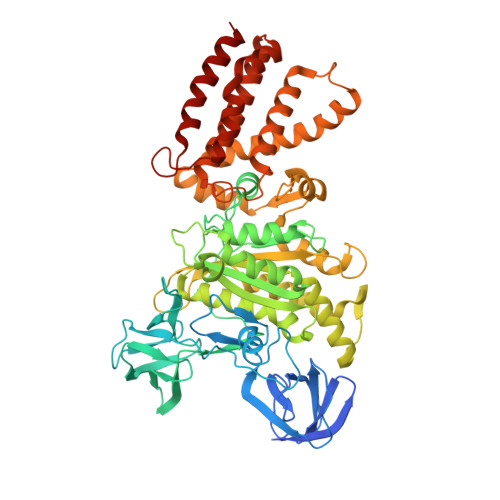
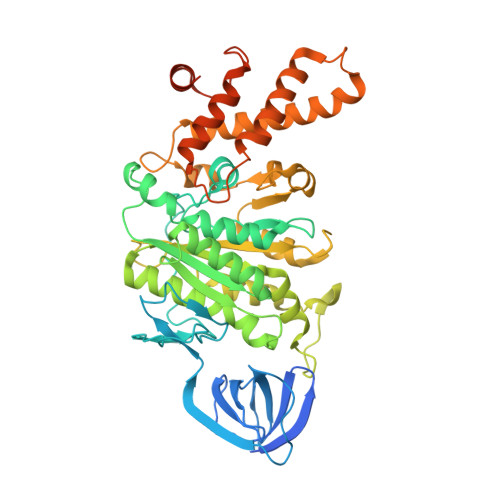
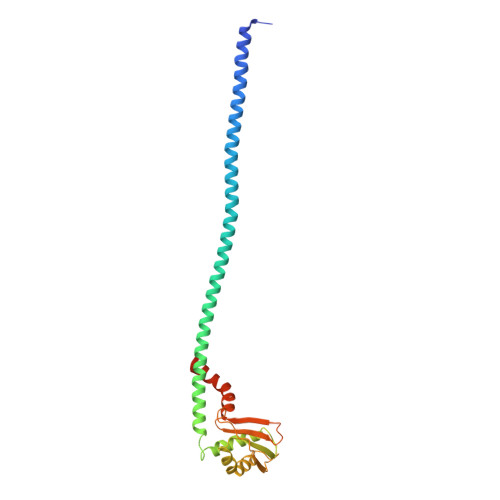
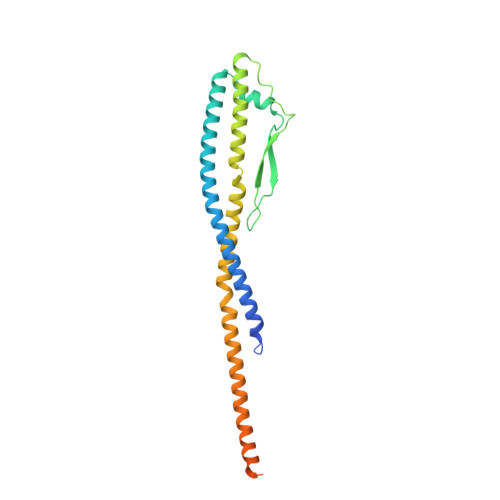
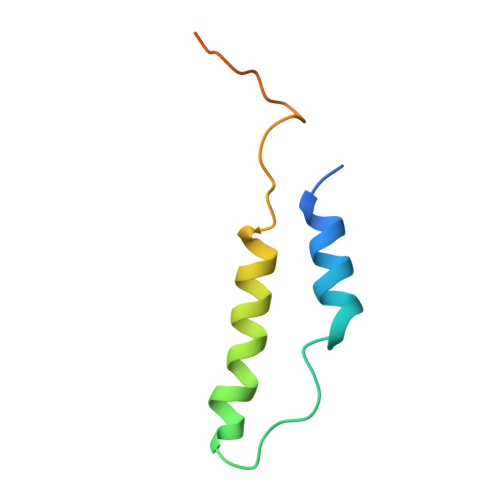
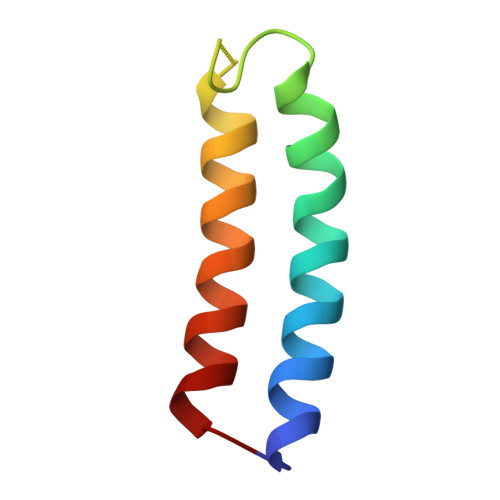
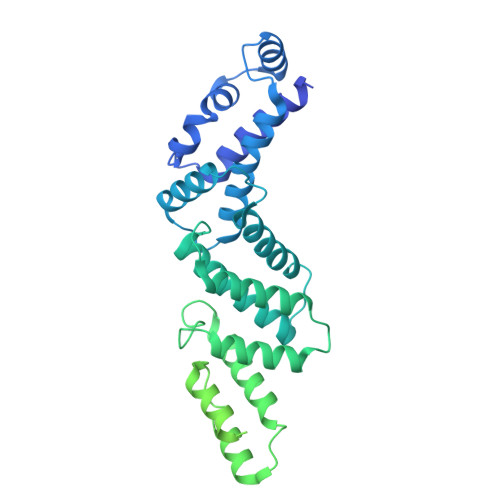
Molecular basis for the binding and modulation of V-ATPase by a bacterial effector protein.
Zhao, J., Beyrakhova, K., Liu, Y., Alvarez, C.P., Bueler, S.A., Xu, L., Xu, C., Boniecki, M.T., Kanelis, V., Luo, Z.Q., Cygler, M., Rubinstein, J.L.(2017) PLoS Pathog 13: e1006394-e1006394
- PubMed: 28570695 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.ppat.1006394
- Primary Citation Related Structures:
5UF5, 5UFK, 5VOX, 5VOY, 5VOZ - PubMed Abstract:
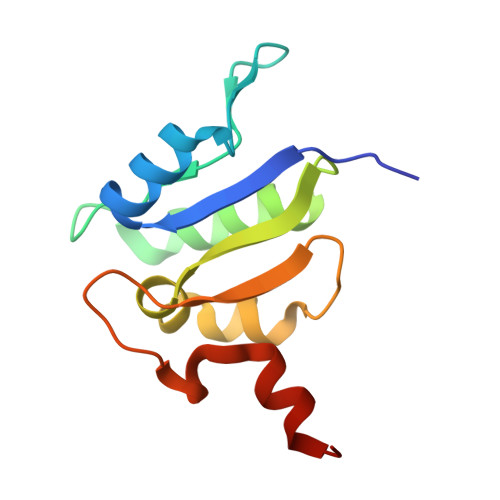
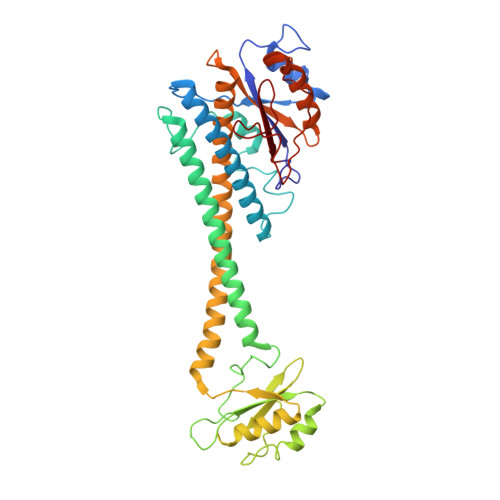
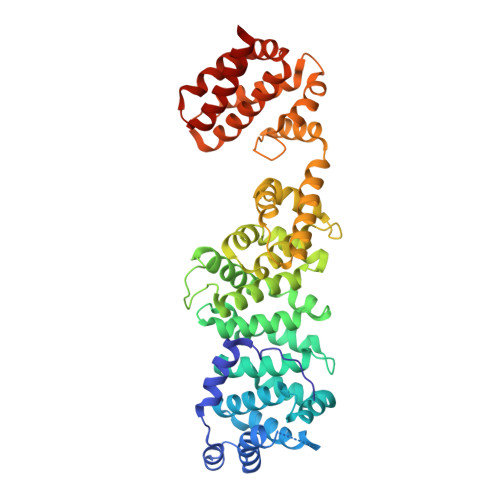
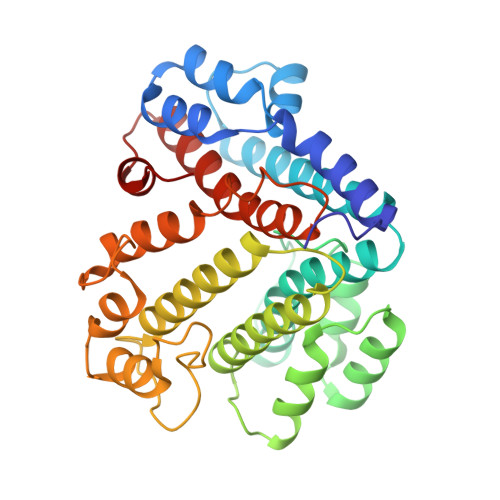
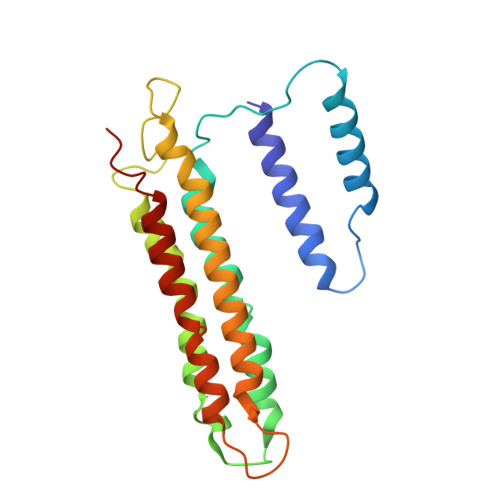
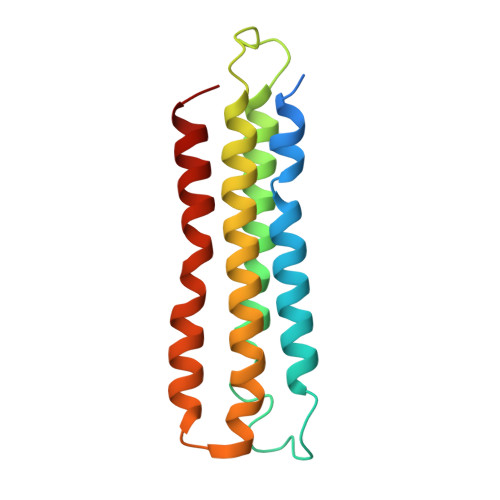
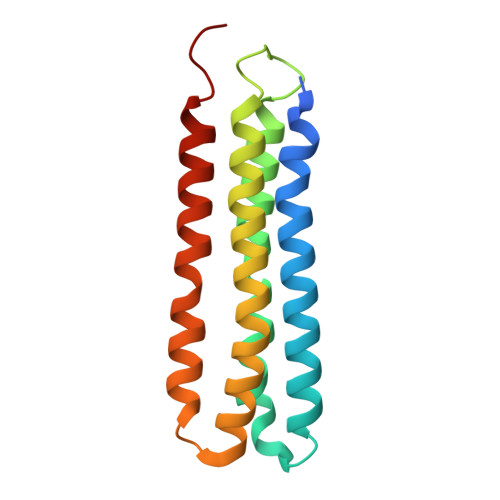
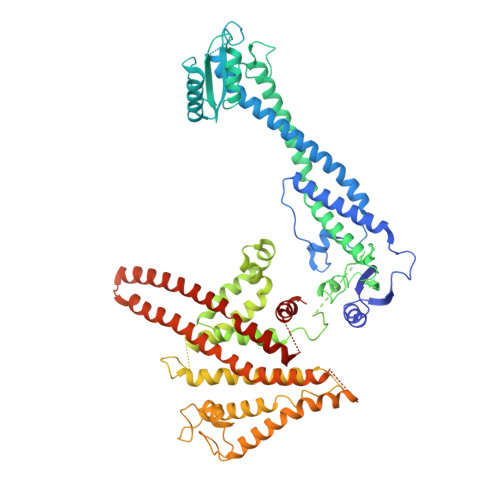
Intracellular pathogenic bacteria evade the immune response by replicating within host cells. Legionella pneumophila, the causative agent of Legionnaires' Disease, makes use of numerous effector proteins to construct a niche supportive of its replication within phagocytic cells. The L. pneumophila effector SidK was identified in a screen for proteins that reduce the activity of the proton pumping vacuolar-type ATPases (V-ATPases) when expressed in the yeast Saccharomyces cerevisae. SidK is secreted by L. pneumophila in the early stages of infection and by binding to and inhibiting the V-ATPase, SidK reduces phagosomal acidification and promotes survival of the bacterium inside macrophages. We determined crystal structures of the N-terminal region of SidK at 2.3 Å resolution and used single particle electron cryomicroscopy (cryo-EM) to determine structures of V-ATPase:SidK complexes at ~6.8 Å resolution. SidK is a flexible and elongated protein composed of an α-helical region that interacts with subunit A of the V-ATPase and a second region of unknown function that is flexibly-tethered to the first. SidK binds V-ATPase strongly by interacting via two α-helical bundles at its N terminus with subunit A. In vitro activity assays show that SidK does not inhibit the V-ATPase completely, but reduces its activity by ~40%, consistent with the partial V-ATPase deficiency phenotype its expression causes in yeast. The cryo-EM analysis shows that SidK reduces the flexibility of the A-subunit that is in the 'open' conformation. Fluorescence experiments indicate that SidK binding decreases the affinity of V-ATPase for a fluorescent analogue of ATP. Together, these results reveal the structural basis for the fine-tuning of V-ATPase activity by SidK.
- The Hospital for Sick Children Research Institute, Toronto, Ontario, Canada.
Organizational Affiliation: