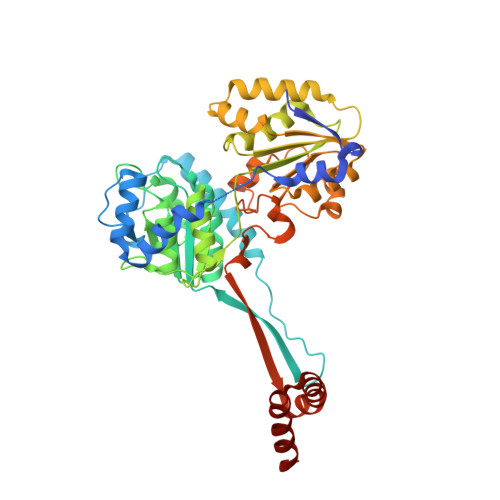
Structures of Medicago truncatula L-Histidinol Dehydrogenase Show Rearrangements Required for NAD(+) Binding and the Cofactor Positioned to Accept a Hydride.
Ruszkowski, M., Dauter, Z.(2017) Sci Rep 7: 10476-10476
- PubMed: 28874718 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-017-10859-0
- Primary Citation Related Structures:
5VLB, 5VLC, 5VLD - PubMed Abstract:
Plants, lower eukaryotes, bacteria, and archaebacteria synthesise L-histidine (His) in a similar, multistep pathway that is absent in mammals. This makes the His biosynthetic route a promising target for herbicides, antifungal agents, and antibiotics. The last enzyme of the pathway, bifunctional L-histidinol dehydrogenase (HDH, EC 1.1.1.23), catalyses two oxidation reactions: from L-histidinol (HOL) to L-histidinaldehyde and from L-histidinaldehyde to His. Over the course of the reaction, HDH utilises two molecules of NAD + as the hydride acceptor. The object of this study was the HDH enzyme from the model legume plant, Medicago truncatula (MtHDH). Three crystal structures complexed with imidazole, HOL, and His with NAD + provided in-depth insights into the enzyme architecture, its active site, and the cofactor binding mode. The overall structure of MtHDH is similar to the two bacterial orthologues whose three-dimensional structures have been determined. The three snapshots, with the MtHDH enzyme captured in different states, visualise structural rearrangements that allow for NAD + binding for the first time. Furthermore, the MtHDH complex with His and NAD + displays the cofactor molecule situated in a way that would allow for a hydride transfer.
- Synchrotron Radiation Research Section of MCL, National Cancer Institute, Argonne, IL, USA. milosz.ruszkowski@nih.gov.
Organizational Affiliation: