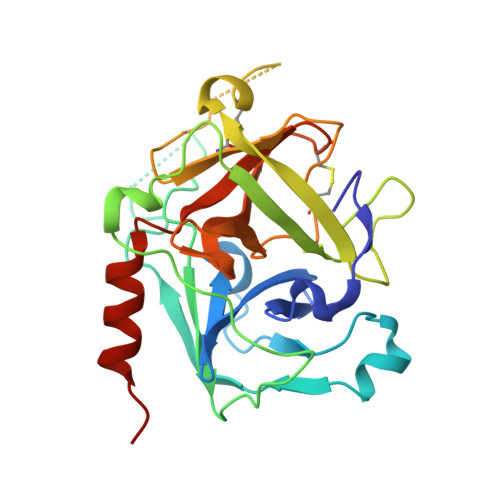
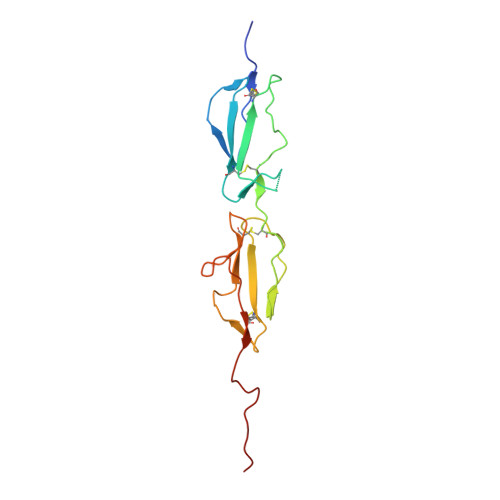
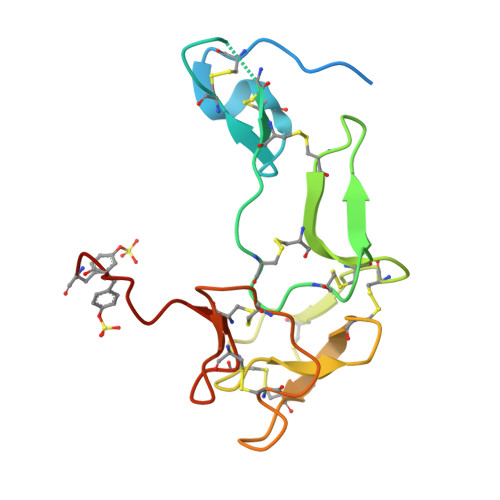
The Structural Basis for Complement Inhibition by Gigastasin, a Protease Inhibitor from the Giant Amazon Leech.
Pang, S.S., Wijeyewickrema, L.C., Hor, L., Tan, S., Lameignere, E., Conway, E.M., Blom, A.M., Mohlin, F.C., Liu, X., Payne, R.J., Whisstock, J.C., Pike, R.N.(2017) J Immunol 199: 3883-3891
- PubMed: 29061764 Search on PubMed
- DOI: https://doi.org/10.4049/jimmunol.1700158
- Primary Citation Related Structures:
5UBM - PubMed Abstract:
Complement is crucial to the immune response, but dysregulation of the system causes inflammatory disease. Complement is activated by three pathways: classical, lectin, and alternative. The classical and lectin pathways are initiated by the C1r/C1s (classical) and MASP-1/MASP-2 (lectin) proteases. Given the role of complement in disease, there is a requirement for inhibitors to control the initiating proteases. In this article, we show that a novel inhibitor, gigastasin, from the giant Amazon leech, potently inhibits C1s and MASP-2, whereas it is also a good inhibitor of MASP-1. Gigastasin is a poor inhibitor of C1r. The inhibitor blocks the active sites of C1s and MASP-2, as well as the anion-binding exosites of the enzymes via sulfotyrosine residues. Complement deposition assays revealed that gigastasin is an effective inhibitor of complement activation in vivo, especially for activation via the lectin pathway. These data suggest that the cumulative effects of inhibiting both MASP-2 and MASP-1 have a greater effect on the lectin pathway than the more potent inhibition of only C1s of the classical pathway.
- Department of Biochemistry and Molecular Biology and Australian Research Council Centre of Excellence in Advanced Molecular Imaging, Biomedicine Discovery Institute, Monash University, Melbourne, Victoria 3800, Australia.
Organizational Affiliation: