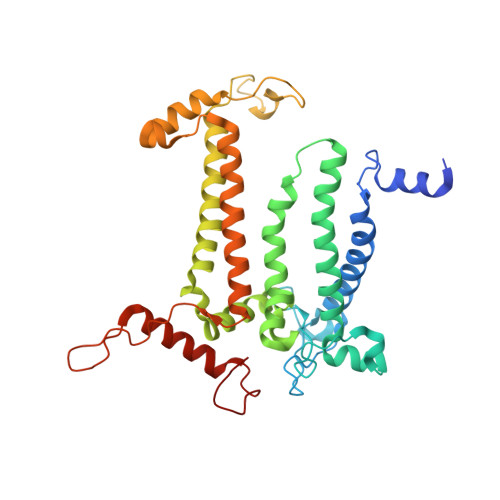
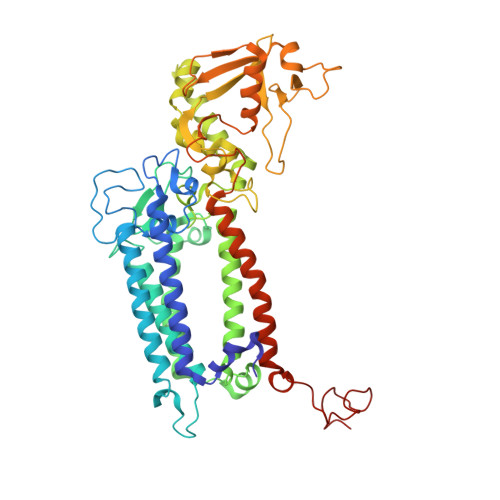
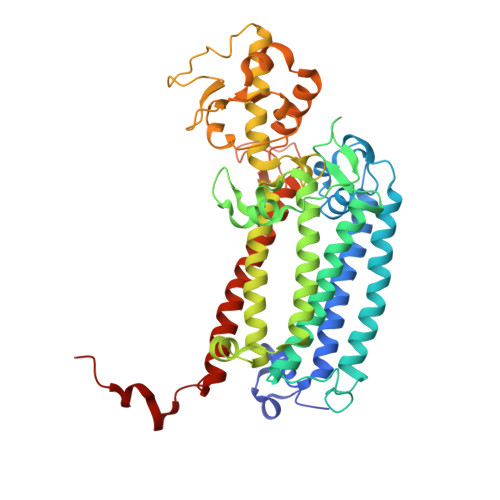
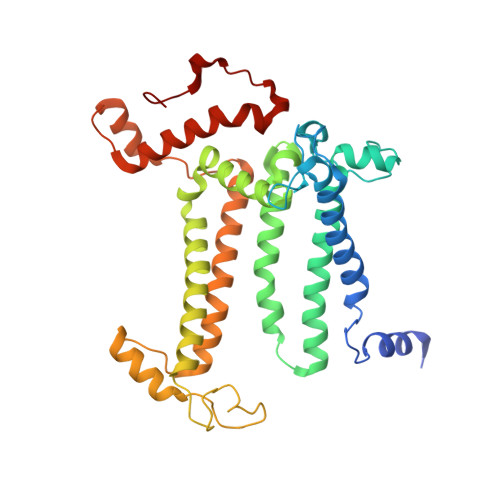
Structure of photosystem II and substrate binding at room temperature.
Young, I.D., Ibrahim, M., Chatterjee, R., Gul, S., Fuller, F.D., Koroidov, S., Brewster, A.S., Tran, R., Alonso-Mori, R., Kroll, T., Michels-Clark, T., Laksmono, H., Sierra, R.G., Stan, C.A., Hussein, R., Zhang, M., Douthit, L., Kubin, M., de Lichtenberg, C., Vo Pham, L., Nilsson, H., Cheah, M.H., Shevela, D., Saracini, C., Bean, M.A., Seuffert, I., Sokaras, D., Weng, T.C., Pastor, E., Weninger, C., Fransson, T., Lassalle, L., Brauer, P., Aller, P., Docker, P.T., Andi, B., Orville, A.M., Glownia, J.M., Nelson, S., Sikorski, M., Zhu, D., Hunter, M.S., Lane, T.J., Aquila, A., Koglin, J.E., Robinson, J., Liang, M., Boutet, S., Lyubimov, A.Y., Uervirojnangkoorn, M., Moriarty, N.W., Liebschner, D., Afonine, P.V., Waterman, D.G., Evans, G., Wernet, P., Dobbek, H., Weis, W.I., Brunger, A.T., Zwart, P.H., Adams, P.D., Zouni, A., Messinger, J., Bergmann, U., Sauter, N.K., Kern, J., Yachandra, V.K., Yano, J.(2016) Nature 540: 453-457
- PubMed: 27871088 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature20161
- Primary Citation Related Structures:
5KAF, 5KAI, 5TIS - PubMed Abstract:
Light-induced oxidation of water by photosystem II (PS II) in plants, algae and cyanobacteria has generated most of the dioxygen in the atmosphere. PS II, a membrane-bound multi-subunit pigment protein complex, couples the one-electron photochemistry at the reaction centre with the four-electron redox chemistry of water oxidation at the Mn 4 CaO 5 cluster in the oxygen-evolving complex (OEC). Under illumination, the OEC cycles through five intermediate S-states (S 0 to S 4 ), in which S 1 is the dark-stable state and S 3 is the last semi-stable state before O-O bond formation and O 2 evolution. A detailed understanding of the O-O bond formation mechanism remains a challenge, and will require elucidation of both the structures of the OEC in the different S-states and the binding of the two substrate waters to the catalytic site. Here we report the use of femtosecond pulses from an X-ray free electron laser (XFEL) to obtain damage-free, room temperature structures of dark-adapted (S 1 ), two-flash illuminated (2F; S 3 -enriched), and ammonia-bound two-flash illuminated (2F-NH 3 ; S 3 -enriched) PS II. Although the recent 1.95 Å resolution structure of PS II at cryogenic temperature using an XFEL provided a damage-free view of the S 1 state, measurements at room temperature are required to study the structural landscape of proteins under functional conditions, and also for in situ advancement of the S-states. To investigate the water-binding site(s), ammonia, a water analogue, has been used as a marker, as it binds to the Mn 4 CaO 5 cluster in the S 2 and S 3 states. Since the ammonia-bound OEC is active, the ammonia-binding Mn site is not a substrate water site. This approach, together with a comparison of the native dark and 2F states, is used to discriminate between proposed O-O bond formation mechanisms.
- Molecular Biophysics and Integrated Bioimaging Division, Lawrence Berkeley National Laboratory, Berkeley, CA 94720, USA.
Organizational Affiliation: