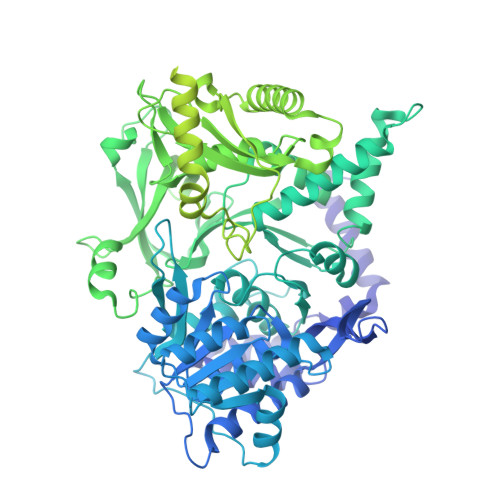
Structures of carboxylic acid reductase reveal domain dynamics underlying catalysis.
Gahloth, D., Dunstan, M.S., Quaglia, D., Klumbys, E., Lockhart-Cairns, M.P., Hill, A.M., Derrington, S.R., Scrutton, N.S., Turner, N.J., Leys, D.(2017) Nat Chem Biol 13: 975-981
- PubMed: 28719588
- DOI: https://doi.org/10.1038/nchembio.2434
- Primary Citation of Related Structures:
5MSC, 5MSD, 5MSO, 5MSP, 5MSR, 5MSS, 5MST, 5MSU, 5MSV, 5MSW - PubMed Abstract:
Carboxylic acid reductase (CAR) catalyzes the ATP- and NADPH-dependent reduction of carboxylic acids to the corresponding aldehydes. The enzyme is related to the nonribosomal peptide synthetases, consisting of an adenylation domain fused via a peptidyl carrier protein (PCP) to a reductase termination domain. Crystal structures of the CAR adenylation-PCP didomain demonstrate that large-scale domain motions occur between the adenylation and thiolation states. Crystal structures of the PCP-reductase didomain reveal that phosphopantetheine binding alters the orientation of a key Asp, resulting in a productive orientation of the bound nicotinamide. This ensures that further reduction of the aldehyde product does not occur. Combining crystallography with small-angle X-ray scattering (SAXS), we propose that molecular interactions between initiation and termination domains are limited to competing PCP docking sites. This theory is supported by the fact that (R)-pantetheine can support CAR activity for mixtures of the isolated domains. Our model suggests directions for further development of CAR as a biocatalyst.
- Manchester Institute of Biotechnology, School of Chemistry, University of Manchester, Manchester, UK.
Organizational Affiliation: