Structure Based Design of Non-Natural Peptidic Macrocyclic Mcl-1 Inhibitors.
Johannes, J.W., Bates, S., Beigie, C., Belmonte, M.A., Breen, J., Cao, S., Centrella, P.A., Clark, M.A., Cuozzo, J.W., Dumelin, C.E., Ferguson, A.D., Habeshian, S., Hargreaves, D., Joubran, C., Kazmirski, S., Keefe, A.D., Lamb, M.L., Lan, H., Li, Y., Ma, H., Mlynarski, S., Packer, M.J., Rawlins, P.B., Robbins, D.W., Shen, H., Sigel, E.A., Soutter, H.H., Su, N., Troast, D.M., Wang, H., Wickson, K.F., Wu, C., Zhang, Y., Zhao, Q., Zheng, X., Hird, A.W.(2017) ACS Med Chem Lett 8: 239-244
- PubMed: 28197319 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.6b00464
- Primary Citation Related Structures:
5KU9, 5MES, 5MEV - PubMed Abstract:
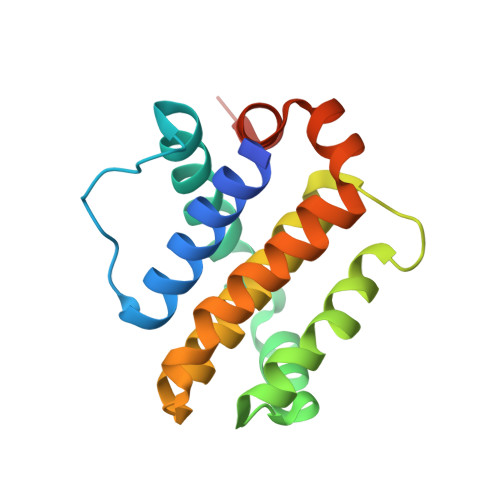
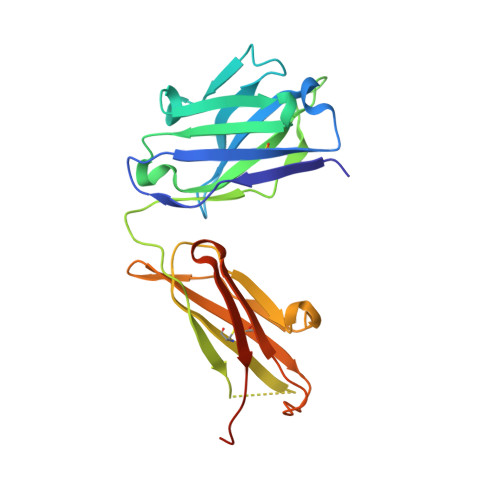
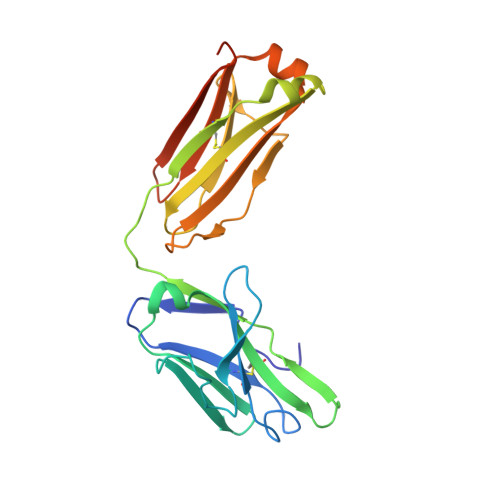
Mcl-1 is a pro-apoptotic BH3 protein family member similar to Bcl-2 and Bcl-xL. Overexpression of Mcl-1 is often seen in various tumors and allows cancer cells to evade apoptosis. Here we report the discovery and optimization of a series of non-natural peptide Mcl-1 inhibitors. Screening of DNA-encoded libraries resulted in hit compound 1 , a 1.5 μM Mcl-1 inhibitor. A subsequent crystal structure demonstrated that compound 1 bound to Mcl-1 in a β-turn conformation, such that the two ends of the peptide were close together. This proximity allowed for the linking of the two ends of the peptide to form a macrocycle. Macrocyclization resulted in an approximately 10-fold improvement in binding potency. Further exploration of a key hydrophobic interaction with Mcl-1 protein and also with the moiety that engages Arg256 led to additional potency improvements. The use of protein-ligand crystal structures and binding kinetics contributed to the design and understanding of the potency gains. Optimized compound 26 is a <3 nM Mcl-1 inhibitor, while inhibiting Bcl-2 at only 5 μM and Bcl-xL at >99 μM, and induces cleaved caspase-3 in MV4-11 cells with an IC 50 of 3 μM after 6 h.
- AstraZeneca R&D Boston , 35 Gatehouse Drive, Waltham, Massachusetts 02451, United States.
Organizational Affiliation: