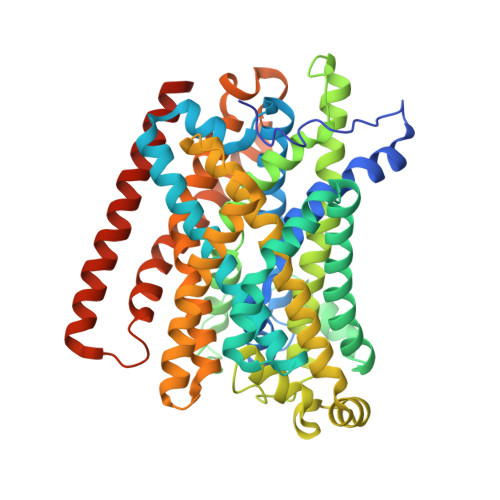
Structural and mechanistic basis of proton-coupled metal ion transport in the SLC11/NRAMP family.
Ehrnstorfer, I.A., Manatschal, C., Arnold, F.M., Laederach, J., Dutzler, R.(2017) Nat Commun 8: 14033-14033
- PubMed: 28059071 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms14033
- Primary Citation Related Structures:
5M87, 5M8A, 5M8J, 5M8K - PubMed Abstract:
Secondary active transporters of the SLC11/NRAMP family catalyse the uptake of iron and manganese into cells. These proteins are highly conserved across all kingdoms of life and thus likely share a common transport mechanism. Here we describe the structural and functional properties of the prokaryotic SLC11 transporter EcoDMT. Its crystal structure reveals a previously unknown outward-facing state of the protein family. In proteoliposomes EcoDMT mediates proton-coupled uptake of manganese at low micromolar concentrations. Mutants of residues in the transition-metal ion-binding site severely affect transport, whereas a mutation of a conserved histidine located near this site results in metal ion transport that appears uncoupled to proton transport. Combined with previous results, our study defines the conformational changes underlying transition-metal ion transport in the SLC11 family and it provides molecular insight to its coupling to protons.
- Department of Biochemistry, University of Zurich, Winterthurerstrasse 190, 8057 Zurich, Switzerland.
Organizational Affiliation: