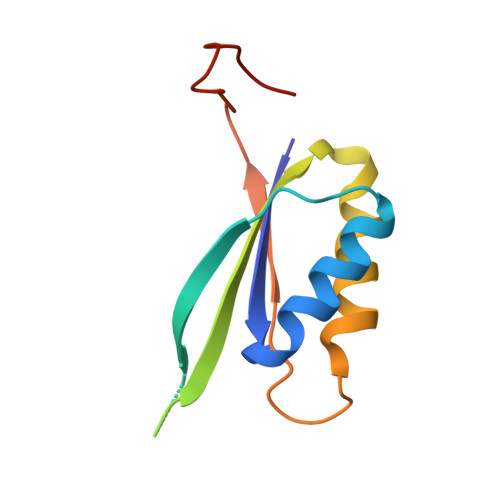
Effects of T-loop modification on the PII-signalling protein: structure of uridylylated Escherichia coli GlnB bound to ATP.
Palanca, C., Rubio, V.(2017) Environ Microbiol Rep 9: 290-299
- PubMed: 28345298 Search on PubMed
- DOI: https://doi.org/10.1111/1758-2229.12533
- Primary Citation Related Structures:
5L9N - PubMed Abstract:
To adapt to environments with variable nitrogen sources and richness, the widely distributed homotrimeric PII signalling proteins bind their allosteric effectors ADP/ATP/2-oxoglutarate, and experience nitrogen-sensitive uridylylation of their flexible T-loops at Tyr51, regulating their interactions with effector proteins. To clarify whether uridylylation triggers a given T-loop conformation, we determined the crystal structure of the classical paradigm of PII protein, Escherichia coli GlnB (EcGlnB), in fully uridylylated form (EcGlnB-UMP 3 ). This is the first structure of a postranslationally modified PII protein. This required recombinant production and purification of the uridylylating enzyme GlnD and its use for full uridylylation of large amounts of recombinantly produced pure EcGlnB. Unlike crystalline non-uridylylated EcGlnB, in which T-loops are fixed, uridylylation rendered the T-loop highly mobile because of loss of contacts mediated by Tyr51, with concomitant abolition of T-loop anchoring via Arg38 on the ATP site. This site was occupied by ATP, providing the first, long-sought snapshot of the EcGlnB-ATP complex, connecting ATP binding with T-loop changes. Inferences are made on the mechanisms of PII selectivity for ATP and of PII-UMP 3 signalling, proposing a model for the architecture of the complex of EcGlnB-UMP 3 with the uridylylation-sensitive PII target ATase (which adenylylates/deadenylylates glutamine synthetase [GS]) and with GS.
- Instituto de Biomedicina de Valencia of the CSIC (IBV-CSIC), Spain.
Organizational Affiliation: