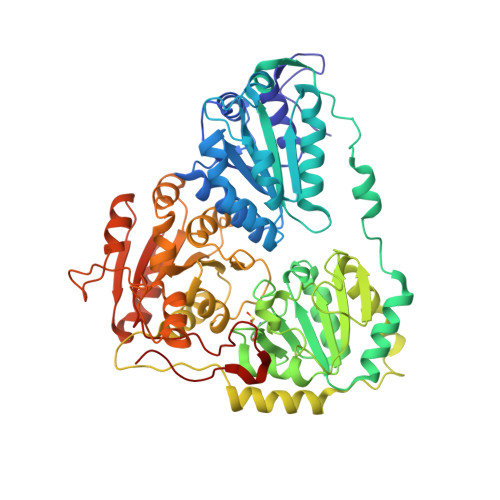
Crystal structure of plant acetohydroxyacid synthase, the target for several commercial herbicides.
Garcia, M.D., Wang, J.G., Lonhienne, T., Guddat, L.W.(2017) FEBS J 284: 2037-2051
- PubMed: 28485824 Search on PubMed
- DOI: https://doi.org/10.1111/febs.14102
- Primary Citation Related Structures:
5K6Q - PubMed Abstract:
Acetohydroxyacid synthase (AHAS, EC 2.2.1.6) is the first enzyme in the branched-chain amino acid biosynthesis pathway. Five of the most widely used commercial herbicides (i.e. sulfonylureas, imidazolinones, triazolopyrimidines, pyrimidinyl-benzoates and sulfonylamino-cabonyl-triazolinones) target this enzyme. Here we have determined the first crystal structure of a plant AHAS in the absence of any inhibitor (2.9 Å resolution) and it shows that the herbicide-binding site adopts a folded state even in the absence of an inhibitor. This is unexpected because the equivalent regions for herbicide binding in uninhibited Saccharomyces cerevisiae AHAS crystal structures are either disordered, or adopt a different fold when the herbicide is not present. In addition, the structure provides an explanation as to why some herbicides are more potent inhibitors of Arabidopsis thaliana AHAS compared to AHASs from other species (e.g. S. cerevisiae). The elucidation of the native structure of plant AHAS provides a new platform for future rational structure-based herbicide design efforts. The coordinates and structure factors for uninhibited AtAHAS have been deposited in the Protein Data Bank (www.pdb.org) with the PDB ID code 5K6Q.
- School of Chemistry and Molecular Biosciences, The University of Queensland, Brisbane, Australia.
Organizational Affiliation: