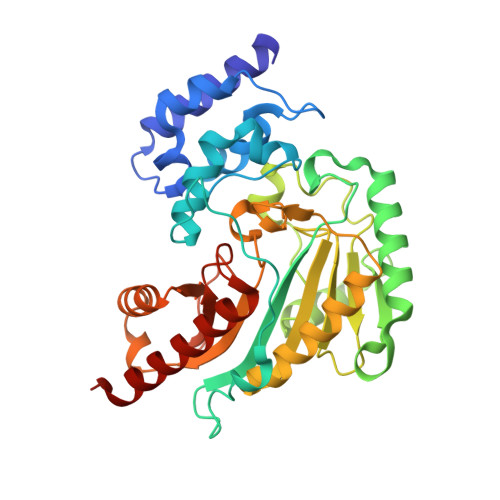
The Sampling of Conformational Dynamics in Ambient-Temperature Crystal Structures of Arginine Kinase.
Godsey, M.H., Davulcu, O., Nix, J.C., Skalicky, J.J., Bruschweiler, R.P., Chapman, M.S.(2016) Structure 24: 1658-1667
- PubMed: 27594681
- DOI: https://doi.org/10.1016/j.str.2016.07.013
- Primary Citation Related Structures:
5J99, 5J9A - PubMed Abstract:
Arginine kinase provides a model for functional dynamics, studied through crystallography, enzymology, and nuclear magnetic resonance. Structures are now solved, at ambient temperature, for the transition state analog (TSA) complex. Analysis of quasi-rigid sub-domain displacements show that differences between the two TSA structures average about 5% of changes between substrate-free and TSA forms, and they are nearly co-linear. Small backbone hinge rotations map to sites that also flex on substrate binding. Anisotropic atomic displacement parameters (ADPs) are refined using rigid-body TLS constraints. Consistency between crystal forms shows that they reflect intrinsic molecular properties more than crystal lattice effects. In many regions, the favored directions of thermal/static displacement are appreciably correlated with movements on substrate binding. Correlation between ADPs and larger substrate-associated movements implies that the latter approximately follow paths of low-energy intrinsic motions.
- Department of Math/Science, Concordia University, Portland, OR 97211, USA.
Organizational Affiliation: