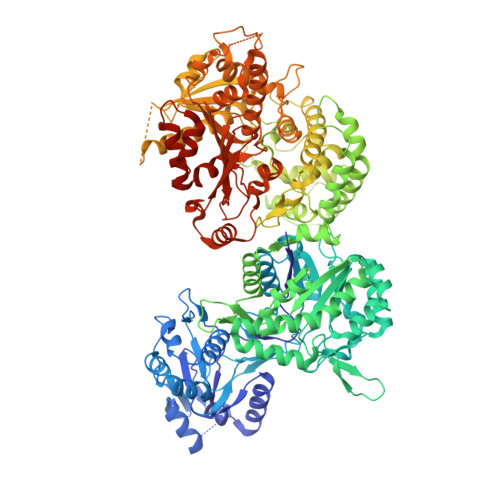
Structural Insights into an Oxalate-producing Serine Hydrolase with an Unusual Oxyanion Hole and Additional Lyase Activity
Oh, J., Hwang, I., Rhee, S.(2016) J Biological Chem 291: 15185-15195
- PubMed: 27226606
- DOI: https://doi.org/10.1074/jbc.M116.727180
- Primary Citation of Related Structures:
5IKY, 5IKZ - PubMed Abstract:
In Burkholderia species, the production of oxalate, an acidic molecule, is a key event for bacterial growth in the stationary phase. Oxalate plays a central role in maintaining environmental pH, which counteracts inevitable population-collapsing alkaline toxicity in amino acid-based culture medium. In the phytopathogen Burkholderia glumae, two enzymes are responsible for oxalate production. First, the enzyme oxalate biosynthetic component A (ObcA) catalyzes the formation of a tetrahedral C6-CoA adduct from the substrates acetyl-CoA and oxaloacetate. Then the ObcB enzyme liberates three products from the C6-CoA adduct: oxalate, acetoacetate, and CoA. Interestingly, these two stepwise reactions are catalyzed by a single bifunctional enzyme, Obc1, from Burkholderia thailandensis and Burkholderia pseudomallei Obc1 has an ObcA-like N-terminal domain and shows ObcB activity in its C-terminal domain despite no sequence homology with ObcB. We report the crystal structure of Obc1 in its apo and glycerol-bound form at 2.5 Å and 2.8 Å resolution, respectively. The Obc1 N-terminal domain is essentially identical both in structure and function to that of ObcA. Its C-terminal domain has an α/β hydrolase fold that has a catalytic triad for oxalate production and a novel oxyanion hole distinct from the canonical HGGG motif in other α/β hydrolases. Functional analyses through mutagenesis studies suggested that His-934 is an additional catalytic acid/base for its lyase activity and liberates two additional products, acetoacetate and CoA. These results provide structural and functional insights into bacterial oxalogenesis and an example of divergent evolution of the α/β hydrolase fold, which has both hydrolase and lyase activity.
- From the Department of Agricultural Biotechnology, Seoul National University, Seoul 151-921, Korea.
Organizational Affiliation: