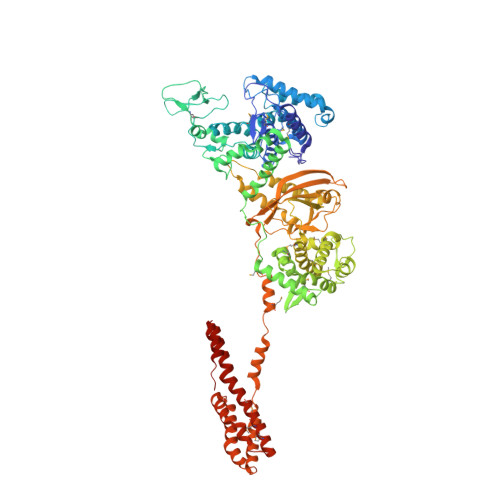
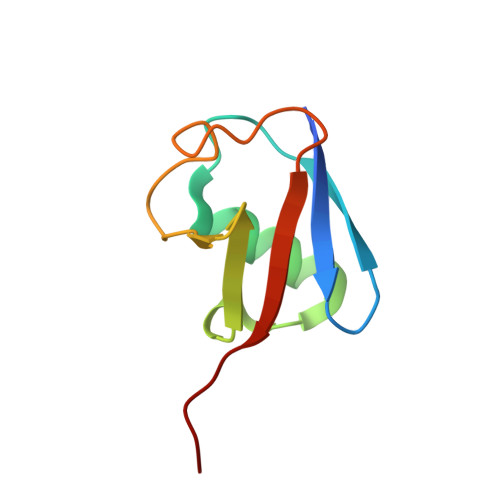
Structural Insights into Non-canonical Ubiquitination Catalyzed by SidE.
Wang, Y., Shi, M., Feng, H., Zhu, Y., Liu, S., Gao, A., Gao, P.(2018) Cell 173: 1231-1243.e16
- PubMed: 29731171 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2018.04.023
- Primary Citation Related Structures:
5ZQ2, 5ZQ3, 5ZQ4, 5ZQ5, 5ZQ6, 5ZQ7 - PubMed Abstract:
Ubiquitination constitutes one of the most important signaling mechanisms in eukaryotes. Conventional ubiquitination is catalyzed by the universally conserved E1-E2-E3 three-enzyme cascade in an ATP-dependent manner. The newly identified SidE family effectors of the pathogen Legionella pneumophila ubiquitinate several human proteins by a different mechanism without engaging any of the conventional ubiquitination machinery. We now report the crystal structures of SidE alone and in complex with ubiquitin, NAD, and ADP-ribose, thereby capturing different conformations of SidE before and after ubiquitin and ligand binding. The structures of ubiquitin bound to both mART and PDE domains reveal several unique features of the two reaction steps catalyzed by SidE. Further, the structural and biochemical results demonstrate that SidE family members do not recognize specific structural folds of the substrate proteins. Our studies provide both structural explanations for the functional observations and new insights into the molecular mechanisms of this non-canonical ubiquitination machinery.
- CAS Key Laboratory of Infection and Immunity, CAS Center for Excellence in Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, China.
Organizational Affiliation: