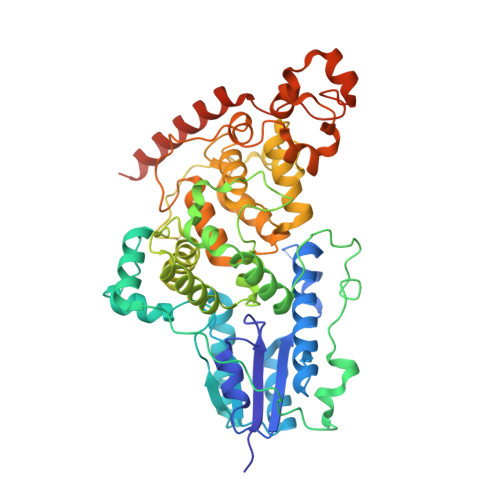
Structure of the bifunctional cryptochrome aCRY from Chlamydomonas reinhardtii
Franz, S., Ignatz, E., Wenzel, S., Zielosko, H., Putu, E.P.G.N., Maestre-Reyna, M., Tsai, M.D., Yamamoto, J., Mittag, M., Essen, L.O.(2018) Nucleic Acids Res 46: 8010-8022
- PubMed: 30032195 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gky621
- Primary Citation Related Structures:
5ZM0, 6FN0, 6FN2, 6FN3 - PubMed Abstract:
Photolyases and cryptochromes form an almost ubiquitous family of blue light photoreceptors involved in the repair and maintenance of DNA integrity or regulatory control. We found that one cryptochrome from the green alga Chlamydomonas reinhardtii (CraCRY) is capable of both, control of transcript levels and the sexual cycle of the alga in a positive (germination) and negative manner (mating ability), as well as catalyzing the repair of UV-DNA lesions. Its 1.6 Å crystal structure shows besides the FAD chromophore an aromatic tetrad that is indispensable in animal-like type I cryptochromes for light-driven change of their signaling-active redox state and formation of a stable radical pair. Given CraCRY's catalytic activity as (6-4) photolyase in vivo and in vitro, we present the first co-crystal structure of a cryptochrome with duplex DNA comprising a (6-4) pyrimidine-pyrimidone lesion. This 2.9 Å structure reveals a distinct conformation for the catalytic histidine His1, H357, that challenges previous models of a single-photon driven (6-4) photolyase mechanism.
- Unit for Structural Biochemistry, Department of Chemistry, Philipps University Marburg, Hans-Meerwein Straße 4, 35032 Marburg, Germany.
Organizational Affiliation: