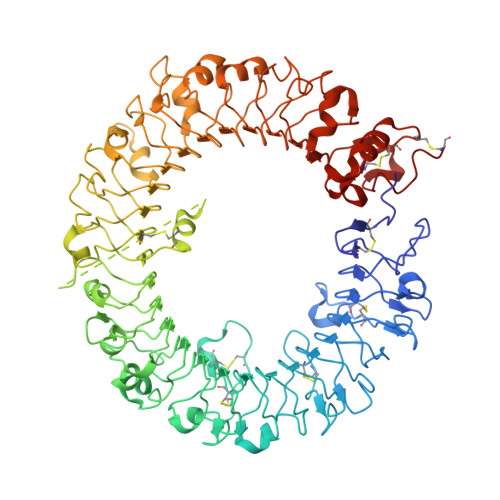
Structural basis for species-specific activation of mouse Toll-like receptor 9
Ishida, H., Ohto, U., Shibata, T., Miyake, K., Shimizu, T.(2018) FEBS Lett 592: 2636-2646
- PubMed: 29961984 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.13176
- Primary Citation Related Structures:
5ZLN - PubMed Abstract:
Single-stranded DNA containing unmethylated cytosine-phosphate-guanine (CpG) motifs derived from microorganisms are recognized by Toll-like receptor (TLR) 9 and activate an innate immune response. TLR9 has two DNA-binding sites for CpG DNA and DNA containing cytosine at the second position from the 5'-end; both are required for efficient TLR9 activation in most vertebrate species. However, mouse TLR9 can be dimerized by CpG DNA only, although the underlying mechanism remains elusive. Here, we report the crystal structure of mouse TLR9 complexed with both DNAs. Although most TLR9-CpG DNA interactions are conserved among species, some are unique to mice and involved in species-specificity. These findings provide the structural basis for how mouse TLR9 dimerizes efficiently in response to CpG DNA to activate innate immunity.
- Graduate School of Pharmaceutical Sciences, The University of Tokyo, Hongo, Bunkyo-ku, Japan.
Organizational Affiliation: