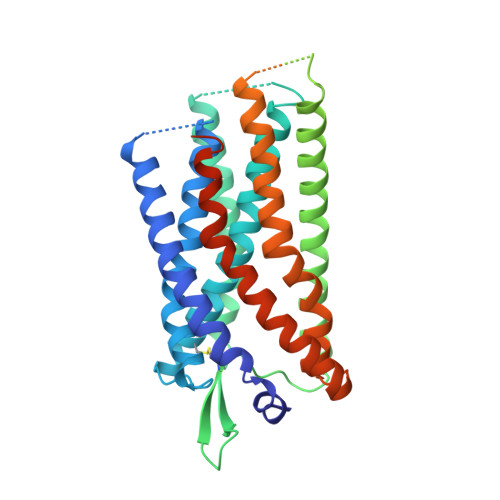
Structural basis for signal recognition and transduction by platelet-activating-factor receptor.
Cao, C., Tan, Q., Xu, C., He, L., Yang, L., Zhou, Y., Zhou, Y., Qiao, A., Lu, M., Yi, C., Han, G.W., Wang, X., Li, X., Yang, H., Rao, Z., Jiang, H., Zhao, Y., Liu, J., Stevens, R.C., Zhao, Q., Zhang, X.C., Wu, B.(2018) Nat Struct Mol Biol 25: 488-495
- PubMed: 29808000 Search on PubMed
- DOI: https://doi.org/10.1038/s41594-018-0068-y
- Primary Citation Related Structures:
5ZKP, 5ZKQ - PubMed Abstract:
Platelet-activating-factor receptor (PAFR) responds to platelet-activating factor (PAF), a phospholipid mediator of cell-to-cell communication that exhibits diverse physiological effects. PAFR is considered an important drug target for treating asthma, inflammation and cardiovascular diseases. Here we report crystal structures of human PAFR in complex with the antagonist SR 27417 and the inverse agonist ABT-491 at 2.8-Å and 2.9-Å resolution, respectively. The structures, supported by molecular docking of PAF, provide insights into the signal-recognition mechanisms of PAFR. The PAFR-SR 27417 structure reveals an unusual conformation showing that the intracellular tips of helices II and IV shift outward by 13 Å and 4 Å, respectively, and helix VIII adopts an inward conformation. The PAFR structures, combined with single-molecule FRET and cell-based functional assays, suggest that the conformational change in the helical bundle is ligand dependent and plays a critical role in PAFR activation, thus greatly extending knowledge about signaling by G-protein-coupled receptors.
- National Laboratory of Biomacromolecules, National Center of Protein Science-Beijing, CAS Center for Excellence in Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing, China.
Organizational Affiliation: