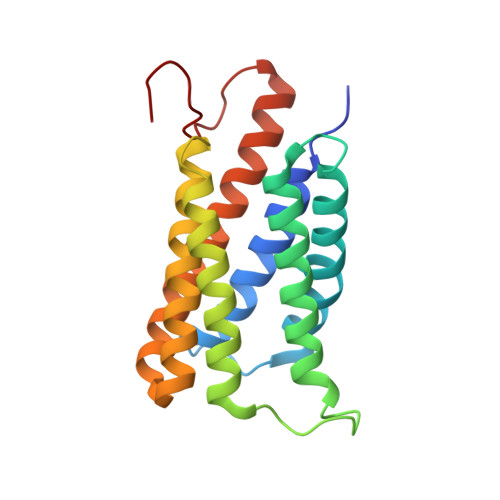
Succinate-acetate permease from Citrobacter koseri is an anion channel that unidirectionally translocates acetate
Qiu, B., Xia, B., Zhou, Q., Lu, Y., He, M., Hasegawa, K., Ma, Z., Zhang, F., Gu, L., Mao, Q., Wang, F., Zhao, S., Gao, Z., Liao, J.(2018) Cell Res 28: 644-654
- PubMed: 29588525 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41422-018-0032-8
- Primary Citation Related Structures:
5YS3, 5YS8 - PubMed Abstract:
Acetate is an important metabolite in metabolism and cell signaling. Succinate-Acetate Permease (SatP) superfamily proteins are known to be responsible for acetate transport across membranes, but the nature of this transport remains unknown. Here, we show that the SatP homolog from Citrobacter koseri (SatP_Ck) is an anion channel that can unidirectionally translocate acetate at rates of the order of ~10 7 ions/s. Crystal structures of SatP_Ck in complex with multiple acetates at 1.8 Å reveal that the acetate pathway consists of four acetate-binding sites aligned in a single file that are interrupted by three hydrophobic constrictions. The bound acetates at the four sites are each orientated differently. The acetate at the cytoplasmic vestibule is partially dehydrated, whereas those in the main pore body are fully dehydrated. Aromatic residues within the substrate pathway may coordinate translocation of acetates via anion-π interactions. SatP_Ck reveals a new type of selective anion channel and provides a structural and functional template for understanding organic anion transport.
- School of Life Science and Technology, ShanghaiTech University, Shanghai, 201210, China.
Organizational Affiliation: