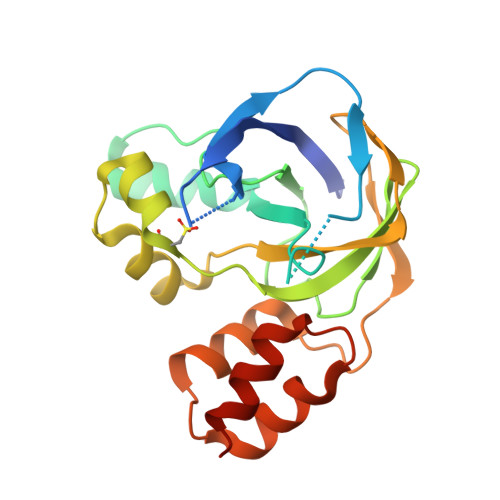
Crystal structure of the hydroxylaminopurine resistance protein, YiiM, and its putative molybdenum cofactor-binding catalytic site.
Namgung, B., Kim, J.H., Song, W.S., Yoon, S.I.(2018) Sci Rep 8: 3304-3304
- PubMed: 29459651 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-018-21660-y
- Primary Citation Related Structures:
5YHH, 5YHI - PubMed Abstract:
The molybdenum cofactor (Moco) is a molybdenum-conjugated prosthetic group that is ubiquitously found in plants, animals, and bacteria. Moco is required for the nitrogen-reducing reaction of the Moco sulfurase C-terminal domain (MOSC) family. Despite the biological significance of MOSC proteins in the conversion of prodrugs and resistance against mutagens, their structural features and Moco-mediated catalysis mechanism have not been described in detail. YiiM is a MOSC protein that is involved in reducing mutagenic 6-N-hydroxylaminopurine to nontoxic adenine in bacteria. Here, we report two crystal structures of YiiM: one from Gram-positive Geobacillus stearothermophilus (gsYiiM) and the other from Gram-negative Escherichia coli (ecYiiM). Although gsYiiM and ecYiiM differ in oligomerization state and protein stability, both consist of three structural modules (a β-barrel and two α-helix bundles) and feature a cavity surrounded by the three modules. The cavity is characterized by positive electrostatic potentials and high sequence conservation. Moreover, the ecYiiM cavity houses a phosphate group, which emulates a part of Moco, and contains a highly reactive invariant cysteine residue. We thus propose that the cavity is the catalytic site where Moco binds and the substrate is reduced. Moreover, our comparative structural analysis highlights the common but distinct structural features of MOSC proteins.
- Division of Biomedical Convergence, College of Biomedical Science, Kangwon National University, Chuncheon, 24341, Republic of Korea.
Organizational Affiliation: