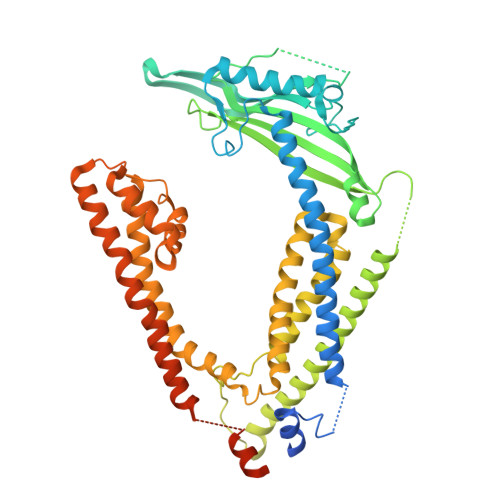
Cryo-EM structures of the mammalian endo-lysosomal TRPML1 channel elucidate the combined regulation mechanism
Zhang, S., Li, N., Zeng, W., Gao, N., Yang, M.(2017) Protein Cell 8: 834-847
- PubMed: 28936784
- DOI: https://doi.org/10.1007/s13238-017-0476-5
- Primary Citation Related Structures:
5YDZ, 5YE1, 5YE2, 5YE5 - PubMed Abstract:
TRPML1 channel is a non-selective group-2 transient receptor potential (TRP) channel with Ca 2+ permeability. Located mainly in late endosome and lysosome of all mammalian cell types, TRPML1 is indispensable in the processes of endocytosis, membrane trafficking, and lysosome biogenesis. Mutations of TRPML1 cause a severe lysosomal storage disorder called mucolipidosis type IV (MLIV). In the present study, we determined the cryo-electron microscopy (cryo-EM) structures of Mus musculus TRPML1 (mTRPML1) in lipid nanodiscs and Amphipols. Two distinct states of mTRPML1 in Amphipols are added to the closed state, on which could represent two different confirmations upon activation and regulation. The polycystin-mucolipin domain (PMD) may sense the luminal/extracellular stimuli and undergo a "move upward" motion during endocytosis, thus triggering the overall conformational change in TRPML1. Based on the structural comparisons, we propose TRPML1 is regulated by pH, Ca 2+ , and phosphoinositides in a combined manner so as to accommodate the dynamic endocytosis process.
- Ministry of Education Key Laboratory of Protein Science, Tsinghua-Peking Joint Center for Life Sciences, Beijing Advanced Innovation Center for Structural Biology, School of Life Sciences, Tsinghua University, Beijing, 100084, China.
Organizational Affiliation: