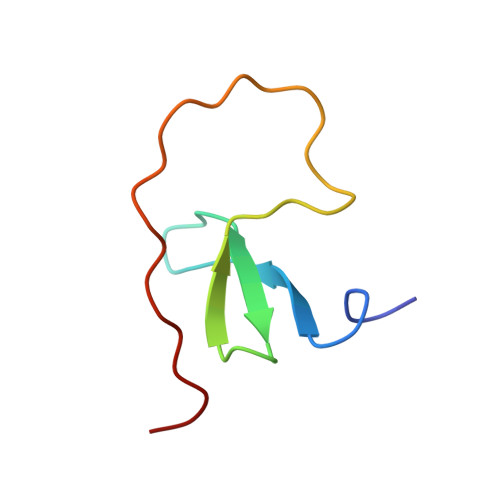
Biophysical studies and NMR structure of YAP2 WW domain - LATS1 PPxY motif complexes reveal the basis of their interaction.
Verma, A., Jing-Song, F., Finch-Edmondson, M.L., Velazquez-Campoy, A., Balasegaran, S., Sudol, M., Sivaraman, J.(2018) Oncotarget 9: 8068-8080
- PubMed: 29487715 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.18632/oncotarget.23909
- Primary Citation Related Structures:
5YDX, 5YDY - PubMed Abstract:
YES-associated protein (YAP) is a major effector protein of the Hippo tumor suppressor pathway, and is phosphorylated by the serine/threonine kinase LATS. Their binding is mediated by the interaction between WW domains of YAP and PPxY motifs of LATS. Their isoforms, YAP2 and LATS1 contain two WW domains and two PPxY motifs respectively. Here, we report the study of the interaction of these domains both in vitro and in human cell lines, to better understand the mechanism of their binding. We show that there is a reciprocal binding preference of YAP2-WW1 with LATS1-PPxY2, and YAP2-WW2 with LATS1-PPxY1. We solved the NMR structures of these complexes and identified several conserved residues that play a critical role in binding. We further created a YAP2 mutant by swapping the WW domains, and found that YAP2 phosphorylation at S127 by LATS1 is not affected by the spatial configuration of its WW domains. This is likely because the region between the PPxY motifs of LATS1 is unstructured, even upon binding with its partner. Based on our observations, we propose possible models for the interaction between YAP2 and LATS1.
- Department of Biological Sciences, National University of Singapore, Singapore.
Organizational Affiliation: