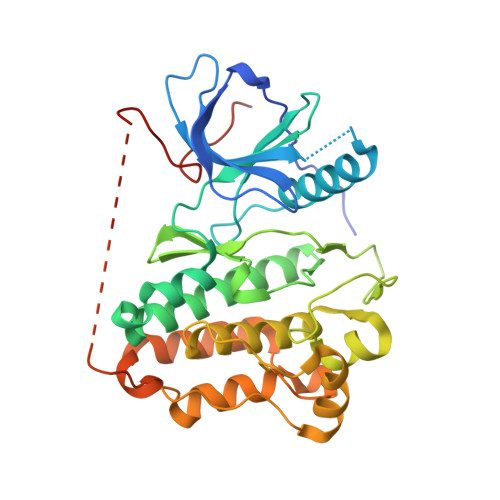
Pharmacological and Structural Characterizations of Naquotinib, a Novel Third-Generation EGFR Tyrosine Kinase Inhibitor, inEGFR-Mutated Non-Small Cell Lung Cancer.
Hirano, T., Yasuda, H., Hamamoto, J., Nukaga, S., Masuzawa, K., Kawada, I., Naoki, K., Niimi, T., Mimasu, S., Sakagami, H., Soejima, K., Betsuyaku, T.(2018) Mol Cancer Ther 17: 740-750
- PubMed: 29467275 Search on PubMed
- DOI: https://doi.org/10.1158/1535-7163.MCT-17-1033
- Primary Citation Related Structures:
5Y9T - PubMed Abstract:
Multiple epidermal growth factor receptor (EGFR) tyrosine kinase inhibitors (EGFR-TKI) have been developed to effectively inhibit EGFR-derived signals in non-small cell lung cancer (NSCLC). In this study, we assessed the efficacy of EGFR-TKIs, including a novel third-generation inhibitor naquotinib (ASP8273), in clinically relevant EGFR mutations, including L858R, exon 19 deletion, L858R+T790M, exon 19 deletion+T790M with or without a C797S mutation, and several exon 20 insertion mutations. Using structural analyses, we also elucidated the mechanism of activation and sensitivity/resistance to EGFR-TKIs in EGFR exon 20 insertion mutations. The efficacy of naquotinib in cells with L858R, exon 19 deletion and exon 19 deletion+T790M was comparable with that of osimertinib. Interestingly, naquotinib was more potent than osimertinib for L858R+T790M. Additionally, naquotinib and osimertinib had comparable efficacy and a wide therapeutic window for cells with EGFR exon 20 insertions. Structural modeling partly elucidated the mechanism of activation and sensitivity/resistance to EGFR-TKIs in two EGFR exon 20 insertion mutants, A767_V769dupASV and Y764_V765insHH. In summary, we have characterized the efficacy of EGFR-TKIs for NSCLC using in vitro and structural analyses and suggested the mechanism of activation and resistance to EGFR-TKIs of EGFR exon 20 insertion mutations. Our findings should guide the selection of appropriate EGFR-TKIs for the treatment of NSCLC with EGFR mutations and help clarify the biology of EGFR exon 20 insertion mutations. Mol Cancer Ther; 17(4); 740-50. ©2018 AACR .
- Division of Pulmonary Medicine, Department of Medicine, Keio University, School of Medicine, Tokyo, Japan.
Organizational Affiliation: