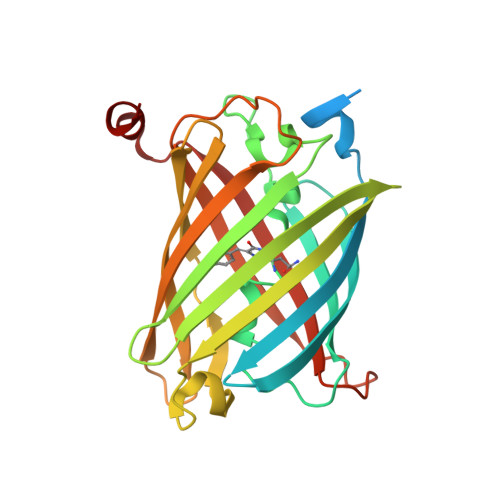
Acid-Tolerant Monomeric GFP from Olindias formosa.
Shinoda, H., Ma, Y., Nakashima, R., Sakurai, K., Matsuda, T., Nagai, T.(2018) Cell Chem Biol 25: 330-338.e7
- PubMed: 29290624 Search on PubMed
- DOI: https://doi.org/10.1016/j.chembiol.2017.12.005
- Primary Citation Related Structures:
5Y00, 5Y01 - PubMed Abstract:
The fluorescent protein (FP) color palette has greatly contributed to the visualization of molecular and cellular processes. However, most FPs lose fluorescence at a pH lower than their neutral pK a (∼6), and this has hampered their application in acidic organelles (pH ∼4.5-6.0). Currently, several cyan- and red-colored acid-tolerant FPs are available; however, there are few reports of acid-tolerant green FPs (GFPs) that are practically applicable to bioimaging. Here, we developed the acid-tolerant monomeric GFP "Gamillus" from the jellyfish Olindias formosa, with excellent brightness, maturation speed, and photostability. Results from X-ray crystallography and point mutagenesis suggest that across a broad pH range the acid tolerance is attributed to stabilization of deprotonation in the chromophore phenyl ring by forming a unique trans configuration. We demonstrate that Gamillus can serve as a molecular tag suitable for imaging in acidic organelles through autophagy-mediated molecular tracking to lysosomes.
- Department of Biotechnology, Graduate School of Engineering, Osaka University, 2-1 Yamadaoka, Suita 565-0871, Japan.
Organizational Affiliation: