Extensive Survey of Antibody Invariant Positions for Efficient Chemical Conjugation Using Expanded Genetic Codes.
Kato, A., Kuratani, M., Yanagisawa, T., Ohtake, K., Hayashi, A., Amano, Y., Kimura, K., Yokoyama, S., Sakamoto, K., Shiraishi, Y.(2017) Bioconjug Chem 28: 2099-2108
- PubMed: 28727448 Search on PubMed
- DOI: https://doi.org/10.1021/acs.bioconjchem.7b00265
- Primary Citation Related Structures:
5XHF, 5XHG - PubMed Abstract:
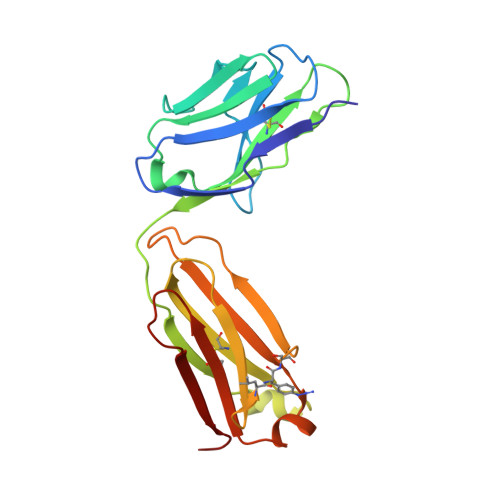
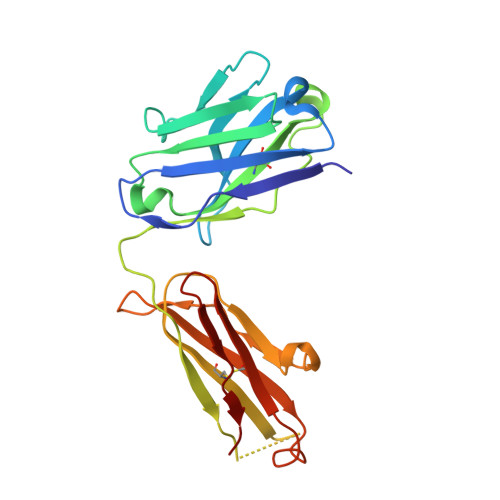
The site-specific chemical conjugation of proteins, following synthesis with an expanded genetic code, promises to advance antibody-based technologies, including antibody drug conjugation and the creation of bispecific Fab dimers. The incorporation of non-natural amino acids into antibodies not only guarantees site specificity but also allows the use of bio-orthogonal chemistry. However, the efficiency of amino acid incorporation fluctuates significantly among different sites, thereby hampering the identification of useful conjugation sites. In this study, we applied the codon reassignment technology to achieve the robust and efficient synthesis of chemically functionalized antibodies containing N ε -(o-azidobenzyloxycarbonyl)-l-lysine (o-Az-Z-Lys) at defined positions. This lysine derivative has a bio-orthogonally reactive group at the end of a long side chain, enabling identification of multiple new positions in Fab-constant domains, allowing chemical conjugation with high efficiency. An X-ray crystallographic study of a Fab variant with o-Az-Z-Lys revealed high-level exposure of the azido group to solvent, with six of the identified positions subsequently used to engineer "Variabodies", a novel antibody format allowing various connections between two Fab molecules. Our findings indicated that some of the created Variabodies exhibited agonistic activity in cultured cells as opposed to the antagonistic nature of antibodies. These results showed that our approach greatly enhanced the availability of antibodies for chemical conjugation and might aid in the development of new therapeutic antibodies.
- RIKEN Structural Biology Laboratory , 1-7-22 Suehiro-cho, Tsurumi-ku, Yokohama 230-0045, Japan.
Organizational Affiliation: