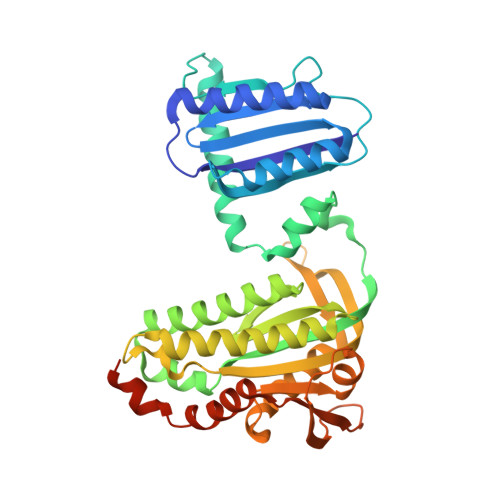
Molecular mechanism of photoactivation of a light-regulated adenylate cyclase.
Ohki, M., Sato-Tomita, A., Matsunaga, S., Iseki, M., Tame, J.R.H., Shibayama, N., Park, S.Y.(2017) Proc Natl Acad Sci U S A 114: 8562-8567
- PubMed: 28739908 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1704391114
- Primary Citation Related Structures:
5X4T, 5X4U, 5X4V - PubMed Abstract:
The photoactivated adenylate cyclase (PAC) from the photosynthetic cyanobacterium Oscillatoria acuminata (OaPAC) detects light through a flavin chromophore within the N-terminal BLUF domain. BLUF domains have been found in a number of different light-activated proteins, but with different relative orientations. The two BLUF domains of OaPAC are found in close contact with each other, forming a coiled coil at their interface. Crystallization does not impede the activity switching of the enzyme, but flash cooling the crystals to cryogenic temperatures prevents the signature spectral changes that occur on photoactivation/deactivation. High-resolution crystallographic analysis of OaPAC in the fully activated state has been achieved by cryocooling the crystals immediately after light exposure. Comparison of the isomorphous light- and dark-state structures shows that the active site undergoes minimal changes, yet enzyme activity may increase up to 50-fold, depending on conditions. The OaPAC models will assist the development of simple, direct means to raise the cyclic AMP levels of living cells by light, and other tools for optogenetics.
- Drug Design Laboratory, Graduate School of Medical Life Science, Yokohama City University, Tsurumi, Yokohama 230-0045, Japan.
Organizational Affiliation: