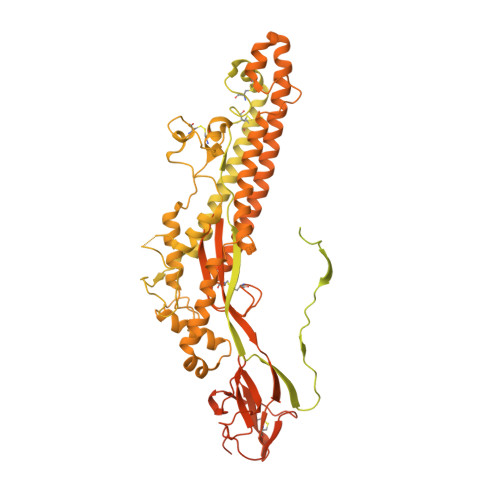
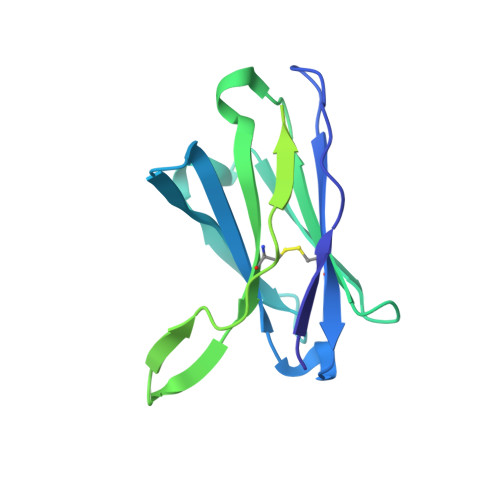
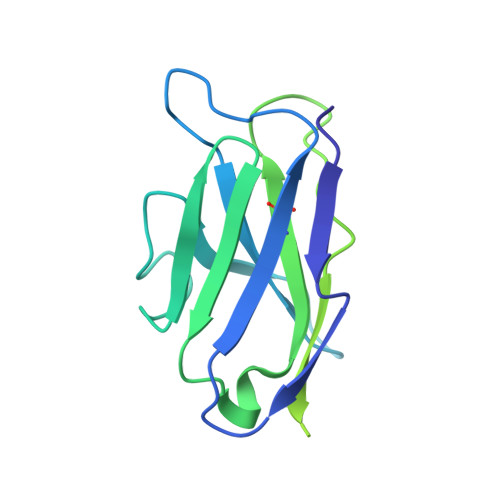
Immunogenicity and structures of a rationally designed prefusion MERS-CoV spike antigen.
Pallesen, J., Wang, N., Corbett, K.S., Wrapp, D., Kirchdoerfer, R.N., Turner, H.L., Cottrell, C.A., Becker, M.M., Wang, L., Shi, W., Kong, W.P., Andres, E.L., Kettenbach, A.N., Denison, M.R., Chappell, J.D., Graham, B.S., Ward, A.B., McLellan, J.S.(2017) Proc Natl Acad Sci U S A 114: E7348-E7357
- PubMed: 28807998
- DOI: https://doi.org/10.1073/pnas.1707304114
- Primary Citation Related Structures:
5VYH, 5VZR, 5W9H, 5W9I, 5W9J, 5W9K, 5W9L, 5W9M, 5W9N, 5W9O, 5W9P - PubMed Abstract:
Middle East respiratory syndrome coronavirus (MERS-CoV) is a lineage C betacoronavirus that since its emergence in 2012 has caused outbreaks in human populations with case-fatality rates of ∼36%. As in other coronaviruses, the spike (S) glycoprotein of MERS-CoV mediates receptor recognition and membrane fusion and is the primary target of the humoral immune response during infection. Here we use structure-based design to develop a generalizable strategy for retaining coronavirus S proteins in the antigenically optimal prefusion conformation and demonstrate that our engineered immunogen is able to elicit high neutralizing antibody titers against MERS-CoV. We also determined high-resolution structures of the trimeric MERS-CoV S ectodomain in complex with G4, a stem-directed neutralizing antibody. The structures reveal that G4 recognizes a glycosylated loop that is variable among coronaviruses and they define four conformational states of the trimer wherein each receptor-binding domain is either tightly packed at the membrane-distal apex or rotated into a receptor-accessible conformation. Our studies suggest a potential mechanism for fusion initiation through sequential receptor-binding events and provide a foundation for the structure-based design of coronavirus vaccines.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA 92037.
Organizational Affiliation: