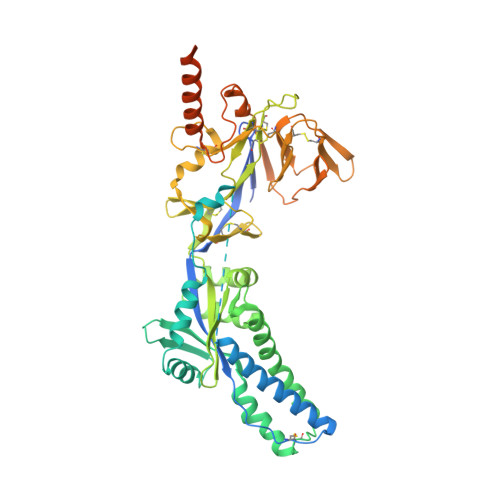
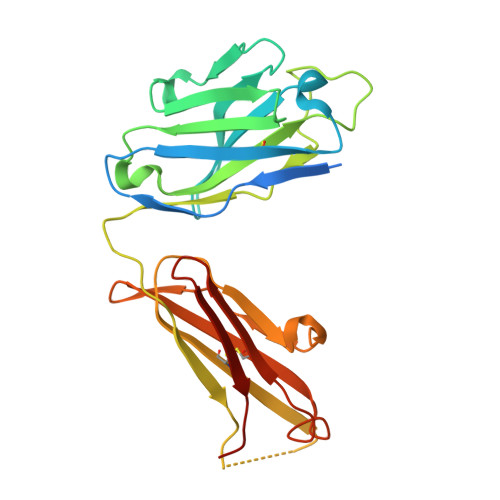
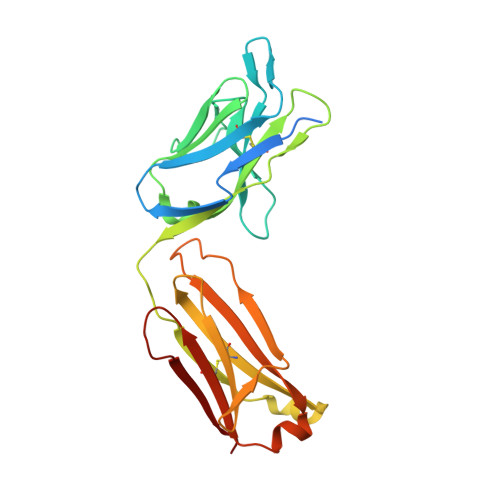
Structural basis of respiratory syncytial virus subtype-dependent neutralization by an antibody targeting the fusion glycoprotein.
Tian, D., Battles, M.B., Moin, S.M., Chen, M., Modjarrad, K., Kumar, A., Kanekiyo, M., Graepel, K.W., Taher, N.M., Hotard, A.L., Moore, M.L., Zhao, M., Zheng, Z.Z., Xia, N.S., McLellan, J.S., Graham, B.S.(2017) Nat Commun 8: 1877-1877
- PubMed: 29187732 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-01858-w
- Primary Citation Related Structures:
5W23, 5W24 - PubMed Abstract:
A licensed vaccine for respiratory syncytial virus (RSV) is unavailable, and passive prophylaxis with the antibody palivizumab is restricted to high-risk infants. Recently isolated antibodies 5C4 and D25 are substantially more potent than palivizumab, and a derivative of D25 is in clinical trials. Here we show that unlike D25, 5C4 preferentially neutralizes subtype A viruses. The crystal structure of 5C4 bound to the RSV fusion (F) protein reveals that the overall binding mode of 5C4 is similar to that of D25, but their angles of approach are substantially different. Mutagenesis and virological studies demonstrate that RSV F residue 201 is largely responsible for the subtype specificity of 5C4. These results improve our understanding of subtype-specific immunity and the neutralization breadth requirements of next-generation antibodies, and thereby contribute to the design of broadly protective RSV vaccines.
- Vaccine Research Center, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD, 20892, USA.
Organizational Affiliation: