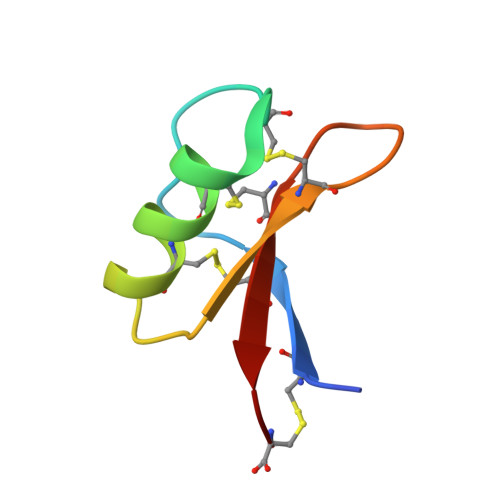
Structure of the defensin NsD7 in complex with PIP2 reveals that defensin : lipid oligomer topologies are dependent on lipid type.
Jarva, M., Lay, F.T., Hulett, M.D., Kvansakul, M.(2017) FEBS Lett 591: 2482-2490
- PubMed: 28741756 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.12761
- Primary Citation Related Structures:
5VYP - PubMed Abstract:
Defensins are innate immune molecules that upon recognition of specific phospholipids can disrupt microbial membranes by forming oligomeric assemblies. Structures of two related plant defensins, NaD1 and NsD7, bound to phosphatidylinositol 4,5-bisphosphate (PIP 2 ) and phosphatidic acid (PA), respectively, revealed striking differences in their oligomeric topologies. To understand how NsD7 binds different phospholipids and rationalize the different topologies, we determined the structure of an NsD7-PIP 2 complex. This structure reveals fundamental differences in phospholipid binding compared to NsD7-PA, and an oligomeric topology nearly identical to the previously determined NaD1-PIP 2 complex, establishing that the PIP 2 fibril topology is conserved between NaD1 and NsD7. Our findings highlight the remarkable ability of defensins to bind different types of phospholipids to form oligomeric fibrils with diverse topologies.
- Department of Biochemistry and Genetics, La Trobe Institute for Molecular Science, La Trobe University, Melbourne, Vic, Australia.
Organizational Affiliation: