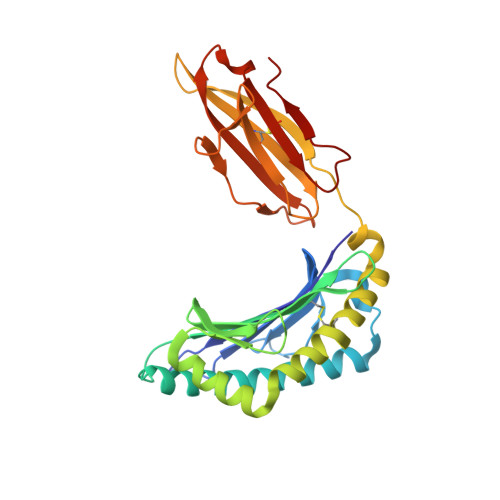
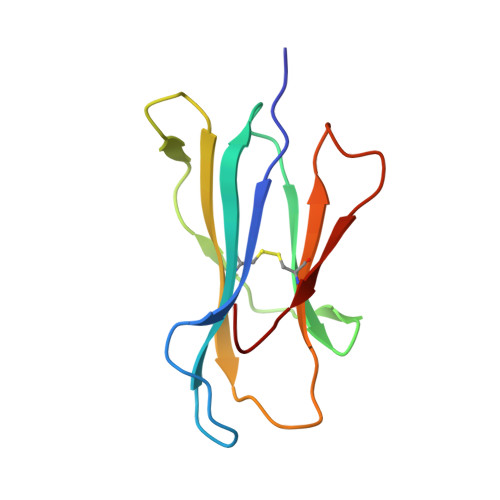
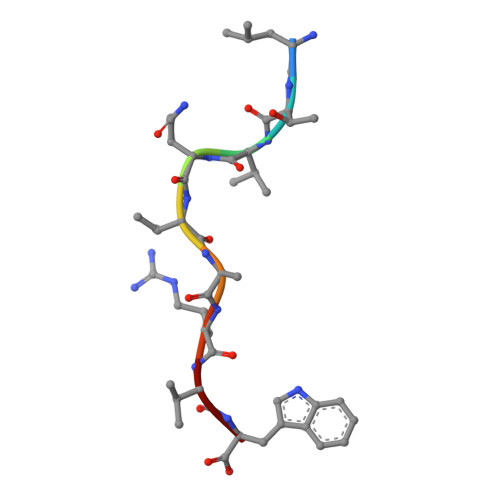
HLA-B57 micropolymorphism defines the sequence and conformational breadth of the immunopeptidome.
Illing, P.T., Pymm, P., Croft, N.P., Hilton, H.G., Jojic, V., Han, A.S., Mendoza, J.L., Mifsud, N.A., Dudek, N.L., McCluskey, J., Parham, P., Rossjohn, J., Vivian, J.P., Purcell, A.W.(2018) Nat Commun 9: 4693-4693
- PubMed: 30410026 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-07109-w
- Primary Citation Related Structures:
5VUD, 5VUE, 5VUF, 5VVP, 5VWD, 5VWF, 5VWH, 5VWJ - PubMed Abstract:
Immunophenotypic differences between closely related human leukocyte antigen (HLA) alleles have been associated with divergent clinical outcomes in infection, autoimmunity, transplantation and drug hypersensitivity. Here we explore the impact of micropolymorphism on peptide antigen presentation by three closely related HLA molecules, HLA-B*57:01, HLA-B*57:03 and HLA-B*58:01, that are differentially associated with the HIV elite controller phenotype and adverse drug reactions. For each allotype, we mine HLA ligand data sets derived from the same parental cell proteome to define qualitative differences in peptide presentation using classical peptide binding motifs and an unbiased statistical approach. The peptide repertoires show marked qualitative overlap, with 982 peptides presented by all allomorphs. However, differences in peptide abundance, HLA-peptide stability, and HLA-bound conformation demonstrate that HLA micropolymorphism impacts more than simply the range of peptide ligands. These differences provide grounds for distinct immune reactivity and insights into the capacity of micropolymorphism to diversify immune outcomes.
- Infection and Immunity Program and Department of Biochemistry and Molecular Biology, Monash Biomedicine Discovery Institute, Monash University, Clayton, VIC, 3800, Australia.
Organizational Affiliation: