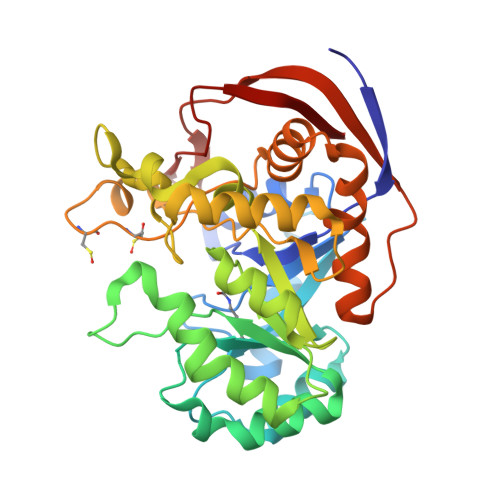
Pyrimidine biosynthesis in pathogens - Structures and analysis of dihydroorotases from Yersinia pestis and Vibrio cholerae.
Lipowska, J., Miks, C.D., Kwon, K., Shuvalova, L., Zheng, H., Lewinski, K., Cooper, D.R., Shabalin, I.G., Minor, W.(2019) Int J Biol Macromol 136: 1176-1187
- PubMed: 31207330 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.ijbiomac.2019.05.149
- Primary Citation Related Structures:
5VGM, 6CTY - PubMed Abstract:
The de novo pyrimidine biosynthesis pathway is essential for the proliferation of many pathogens. One of the pathway enzymes, dihydroorotase (DHO), catalyzes the reversible interconversion of N-carbamoyl-l-aspartate to 4,5-dihydroorotate. The substantial difference between bacterial and mammalian DHOs makes it a promising drug target for disrupting bacterial growth and thus an important candidate to evaluate as a response to antimicrobial resistance on a molecular level. Here, we present two novel three-dimensional structures of DHOs from Yersinia pestis (YpDHO), the plague-causing pathogen, and Vibrio cholerae (VcDHO), the causative agent of cholera. The evaluations of these two structures led to an analysis of all available DHO structures and their classification into known DHO types. Comparison of all the DHO active sites containing ligands that are listed in DrugBank was facilitated by a new interactive, structure-comparison and presentation platform. In addition, we examined the genetic context of characterized DHOs, which revealed characteristic patterns for different types of DHOs. We also generated a homology model for DHO from Plasmodium falciparum.
- Department of Molecular Physiology and Biological Physics, University of Virginia, Charlottesville, VA 22908, USA; Center for Structural Genomics of Infectious Diseases (CSGID), Charlottesville, VA 22908, USA; Faculty of Chemistry, Jagiellonian University, 30-387 Kraków, Poland.
Organizational Affiliation: