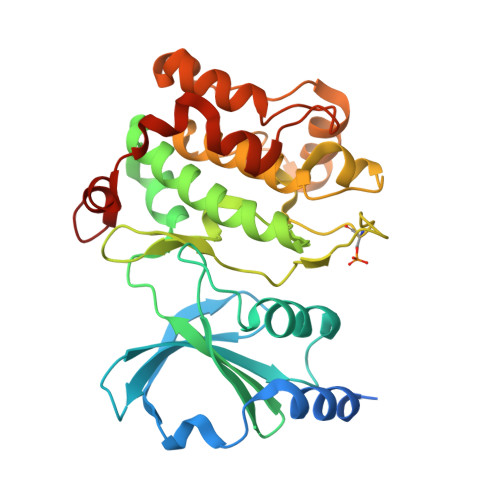
PAK4 crystal structures suggest unusual kinase conformational movements.
Zhang, E.Y., Ha, B.H., Boggon, T.J.(2018) Biochim Biophys Acta 1866: 356-365
- PubMed: 28993291 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bbapap.2017.10.004
- Primary Citation Related Structures:
5VED, 5VEE, 5VEF - PubMed Abstract:
In order for protein kinases to exchange nucleotide they must open and close their catalytic cleft. These motions are associated with rotations of the N-lobe, predominantly around the 'hinge region'. We conducted an analysis of 28 crystal structures of the serine-threonine kinase, p21-activated kinase 4 (PAK4), including three newly determined structures in complex with staurosporine, FRAX486, and fasudil (HA-1077). We find an unusual motion between the N-lobe and C-lobe of PAK4 that manifests as a partial unwinding of helix αC. Principal component analysis of the crystal structures rationalizes these movements into three major states, and analysis of the kinase hydrophobic spines indicates concerted movements that create an accessible back pocket cavity. The conformational changes that we observe for PAK4 differ from previous descriptions of kinase motions, and although we observe these differences in crystal structures there is the possibility that the movements observed may suggest a diversity of kinase conformational changes associated with regulation. Protein kinases are key signaling proteins, and are important drug targets, therefore understanding their regulation is important for both basic research and clinical points of view. In this study, we observe unusual conformational 'hinging' for protein kinases. Hinging, the opening and closing of the kinase sub-domains to allow nucleotide binding and release, is critical for proper kinase regulation and for targeted drug discovery. We determine new crystal structures of PAK4, an important Rho-effector kinase, and conduct analyses of these and previously determined structures. We find that PAK4 crystal structures can be classified into specific conformational groups, and that these groups are associated with previously unobserved hinging motions and an unusual conformation for the kinase hydrophobic core. Our findings therefore indicate that there may be a diversity of kinase hinging motions, and that these may indicate different mechanisms of regulation.
- Department of Molecular Biophysics and Biochemistry, Yale University School of Medicine, 333 Cedar St., New Haven, CT 06520, United States.
Organizational Affiliation: