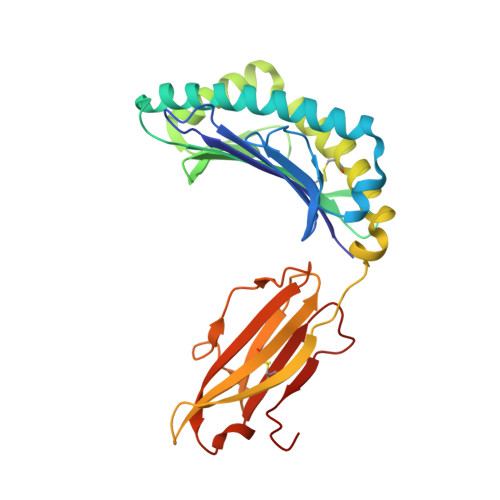
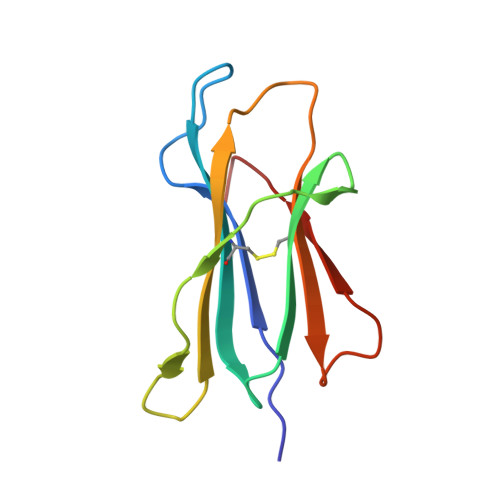
Crystal structure of HLA-B*5801 with a TW10 HIV Gag epitope reveals a novel mode of peptide presentation.
Li, X., Lamothe, P.A., Walker, B.D., Wang, J.H.(2017) Cell Mol Immunol 14: 631-634
- PubMed: 28552904 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/cmi.2017.24
- Primary Citation Related Structures:
5V5L, 5V5M - Department of Medical Oncology and Cancer Biology, Dana-Farber Cancer Institute, Harvard Medical School, Boston, MA 02215, USA.
Organizational Affiliation: