Dominant protection from HLA-linked autoimmunity by antigen-specific regulatory T cells.
Ooi, J.D., Petersen, J., Tan, Y.H., Huynh, M., Willett, Z.J., Ramarathinam, S.H., Eggenhuizen, P.J., Loh, K.L., Watson, K.A., Gan, P.Y., Alikhan, M.A., Dudek, N.L., Handel, A., Hudson, B.G., Fugger, L., Power, D.A., Holt, S.G., Coates, P.T., Gregersen, J.W., Purcell, A.W., Holdsworth, S.R., La Gruta, N.L., Reid, H.H., Rossjohn, J., Kitching, A.R.(2017) Nature 545: 243-247
- PubMed: 28467828 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature22329
- Primary Citation Related Structures:
5V4M, 5V4N - PubMed Abstract:
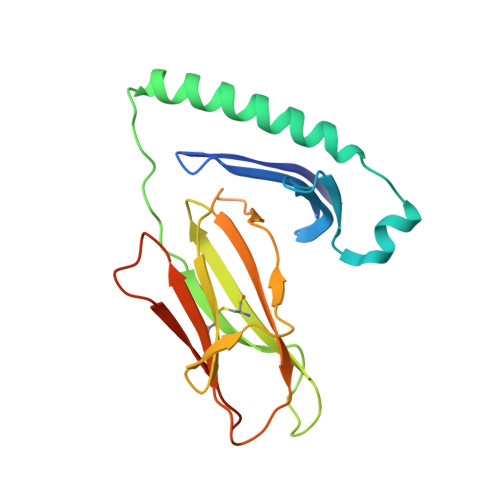
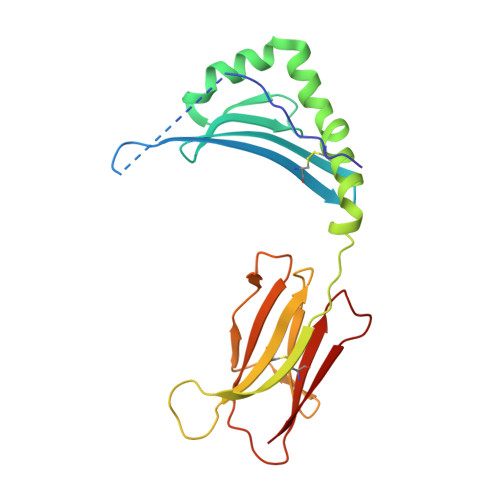
Susceptibility and protection against human autoimmune diseases, including type I diabetes, multiple sclerosis, and Goodpasture disease, is associated with particular human leukocyte antigen (HLA) alleles. However, the mechanisms underpinning such HLA-mediated effects on self-tolerance remain unclear. Here we investigate the molecular mechanism of Goodpasture disease, an HLA-linked autoimmune renal disorder characterized by an immunodominant CD4 + T-cell self-epitope derived from the α3 chain of type IV collagen (α3 135-145 ). While HLA-DR15 confers a markedly increased disease risk, the protective HLA-DR1 allele is dominantly protective in trans with HLA-DR15 (ref. 2). We show that autoreactive α3 135-145 -specific T cells expand in patients with Goodpasture disease and, in α3 135-145 -immunized HLA-DR15 transgenic mice, α3 135-145 -specific T cells infiltrate the kidney and mice develop Goodpasture disease. HLA-DR15 and HLA-DR1 exhibit distinct peptide repertoires and binding preferences and present the α3 135-145 epitope in different binding registers. HLA-DR15-α3 135-145 tetramer + T cells in HLA-DR15 transgenic mice exhibit a conventional T-cell phenotype (T conv ) that secretes pro-inflammatory cytokines. In contrast, HLA-DR1-α3 135-145 tetramer + T cells in HLA-DR1 and HLA-DR15/DR1 transgenic mice are predominantly CD4 + Foxp3 + regulatory T cells (T reg cells) expressing tolerogenic cytokines. HLA-DR1-induced T reg cells confer resistance to disease in HLA-DR15/DR1 transgenic mice. HLA-DR15 + and HLA-DR1 + healthy human donors display altered α3 135-145 -specific T-cell antigen receptor usage, HLA-DR15-α3 135-145 tetramer + Foxp3 - T conv and HLA-DR1-α3 135-145 tetramer + Foxp3 + CD25 hi CD127 lo T reg dominant phenotypes. Moreover, patients with Goodpasture disease display a clonally expanded α3 135-145 -specific CD4 + T-cell repertoire. Accordingly, we provide a mechanistic basis for the dominantly protective effect of HLA in autoimmune disease, whereby HLA polymorphism shapes the relative abundance of self-epitope specific T reg cells that leads to protection or causation of autoimmunity.
- Centre for Inflammatory Diseases, Monash University Department of Medicine, Monash Medical Centre, Clayton, Victoria 3168, Australia.
Organizational Affiliation: