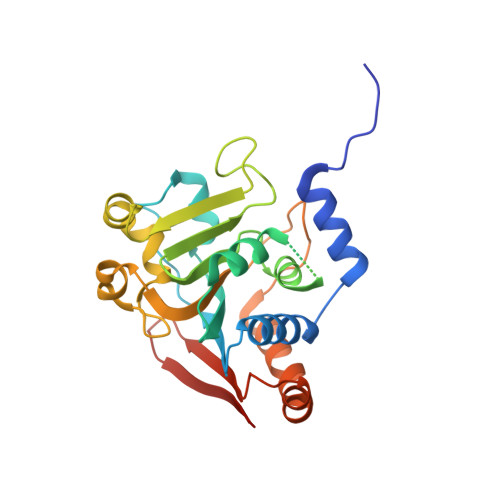
Structures of ubiquitin-like (Ubl) and Hsp90-like domains of sacsin provide insight into pathological mutations.
Menade, M., Kozlov, G., Trempe, J.F., Pande, H., Shenker, S., Wickremasinghe, S., Li, X., Hojjat, H., Dicaire, M.J., Brais, B., McPherson, P.S., Wong, M.J.H., Young, J.C., Gehring, K.(2018) J Biological Chem 293: 12832-12842
- PubMed: 29945973 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA118.003939
- Primary Citation Related Structures:
5V44, 5V45, 5V46, 5V47, 5VSX, 5VSZ - PubMed Abstract:
Autosomal recessive spastic ataxia of Charlevoix-Saguenay (ARSACS) is a neurodegenerative disease that is caused by mutations in the SACS gene. The product of this gene is a very large 520-kDa cytoplasmic protein, sacsin, with a ubiquitin-like (Ubl) domain at the N terminus followed by three large sacsin internal repeat (SIRPT) supradomains and C-terminal J and HEPN domains. The SIRPTs are predicted to contain Hsp90-like domains, suggesting a potential chaperone activity. In this work, we report the structures of the Hsp90-like Sr1 domain of SIRPT1 and the N-terminal Ubl domain determined at 1.55- and 2.1-Å resolutions, respectively. The Ubl domain crystallized as a swapped dimer that could be relevant in the context of full-length protein. The Sr1 domain displays the Bergerat protein fold with a characteristic nucleotide-binding pocket, although it binds nucleotides with very low affinity. The Sr1 structure reveals that ARSACS-causing missense mutations (R272H, R272C, and T201K) disrupt protein folding, most likely leading to sacsin degradation. This work lends structural support to the view of sacsin as a molecular chaperone and provides a framework for future studies of this protein.
- Department of Biochemistry, McGill Centre for Structural Biology, McGill University, Montreal, Quebec H3G 0B1, Canada.
Organizational Affiliation: