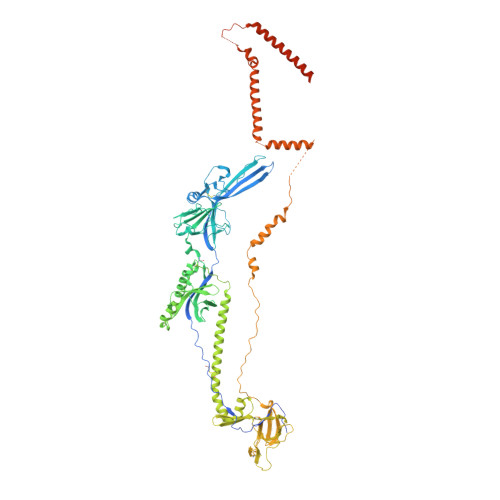
Structural basis for membrane anchoring and fusion regulation of the herpes simplex virus fusogen gB.
Cooper, R.S., Georgieva, E.R., Borbat, P.P., Freed, J.H., Heldwein, E.E.(2018) Nat Struct Mol Biol 25: 416-424
- PubMed: 29728654 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-018-0060-6
- Primary Citation Related Structures:
5V2S, 6BM8 - PubMed Abstract:
Viral fusogens merge viral and cell membranes during cell penetration. Their ectodomains drive fusion by undergoing large-scale refolding, but little is known about the functionally important regions located within or near the membrane. Here we report the crystal structure of full-length glycoprotein B (gB), the fusogen from herpes simplex virus, complemented by electron spin resonance measurements. The membrane-proximal (MPR), transmembrane (TMD), and cytoplasmic (CTD) domains form a uniquely folded trimeric pedestal beneath the ectodomain, which balances dynamic flexibility with extensive, stabilizing membrane interactions. The postfusion conformation of the ectodomain suggests that the CTD likewise adopted the postfusion form. However, hyperfusogenic mutations, which destabilize the prefusion state of gB, target key interfaces and structural motifs that reinforce the observed CTD structure. Thus, a similar CTD structure must stabilize gB in its prefusion state. Our data suggest a model for how this dynamic, membrane-dependent 'clamp' controls the fusogenic refolding of gB.
- Department of Molecular Biology and Microbiology, Tufts University School of Medicine, Boston, MA, USA.
Organizational Affiliation: