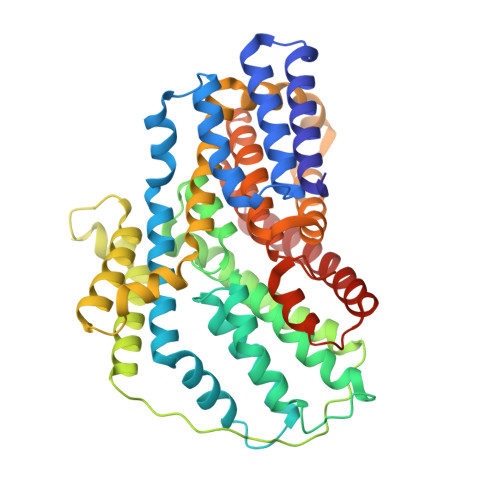
Structure and function of the divalent anion/Na(+) symporter from Vibrio cholerae and a humanized variant.
Nie, R., Stark, S., Symersky, J., Kaplan, R.S., Lu, M.(2017) Nat Commun 8: 15009-15009
- PubMed: 28436435 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms15009
- Primary Citation Related Structures:
5UL7, 5UL9, 5ULD, 5ULE - PubMed Abstract:
Integral membrane proteins of the divalent anion/Na + symporter (DASS) family translocate dicarboxylate, tricarboxylate or sulphate across cell membranes, typically by utilizing the preexisting Na + gradient. The molecular determinants for substrate recognition by DASS remain obscure, largely owing to the absence of any substrate-bound DASS structure. Here we present 2.8-Å resolution X-ray structures of VcINDY, a DASS from Vibrio cholerae that catalyses the co-transport of Na + and succinate. These structures portray the Na + -bound VcINDY in complexes with succinate and citrate, elucidating the binding sites for substrate and two Na + ions. Furthermore, we report the structures of a humanized variant of VcINDY in complexes with succinate and citrate, which predict how a human citrate-transporting DASS may interact with its bound substrate. Our findings provide insights into metabolite transport by DASS, establishing a molecular basis for future studies on the regulation of this transport process.
- Department of Biochemistry and Molecular Biology, Rosalind Franklin University of Medicine and Science, 3333 Green Bay Road, North Chicago, Illinois 60064, USA.
Organizational Affiliation: