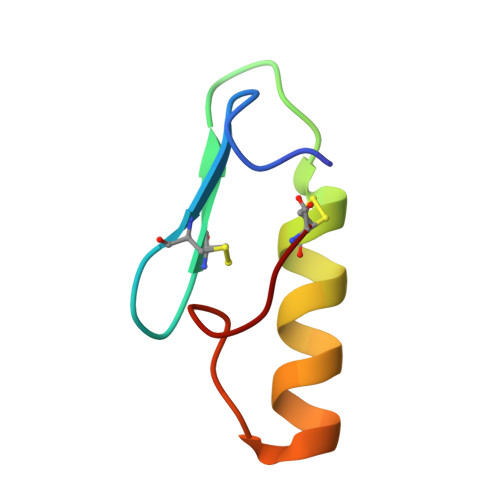
Synthesis, antimicrobial activity and conformational analysis of the class IIa bacteriocin pediocin PA-1 and analogs thereof.
Bedard, F., Hammami, R., Zirah, S., Rebuffat, S., Fliss, I., Biron, E.(2018) Sci Rep 8: 9029-9029
- PubMed: 29899567 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-018-27225-3
- Primary Citation Related Structures:
5UKZ - PubMed Abstract:
The antimicrobial peptide pediocin PA-1 is a class IIa bacteriocin that inhibits several clinically relevant pathogens including Listeria spp. Here we report the synthesis and characterization of whole pediocin PA-1 and novel analogs thereof using a combination of solid- and solution-phase strategies to overcome difficulties due to instability and undesired reactions. Pediocin PA-1 thus synthesized was a potent inhibitor of Listeria monocytogenes (MIC = 6.8 nM), similar to the bacteriocin produced naturally by Pediococcus acidilactici. Of particular interest is that linear analogs lacking both of the disulfide bridges characterizing pediocin PA-1 were as potent. One linear analog was also a strong inhibitor of Clostridium perfringens, another important food-borne pathogen. These results are discussed in light of conformational information derived from circular dichroism, solution NMR spectroscopy and structure-activity relationship studies.
- Faculté de pharmacie, Université Laval and Laboratoire de chimie médicinale, Centre de recherche du CHU de Québec, 2705 Boulevard Laurier, Québec, Québec, G1V 0A6, Canada.
Organizational Affiliation: