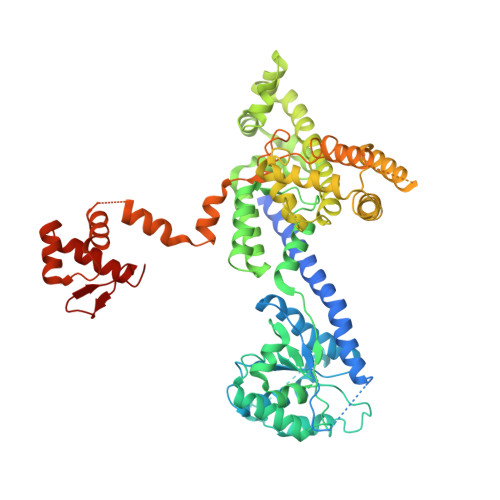
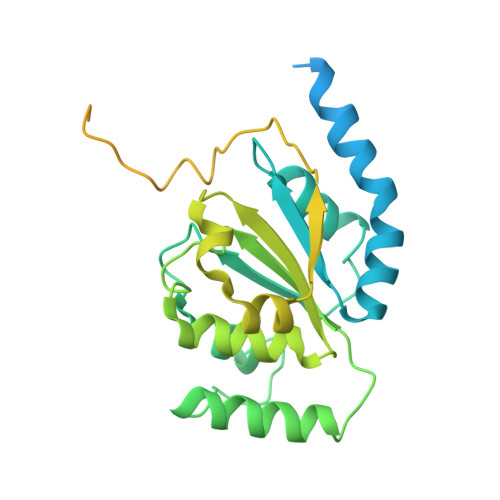
Structure of the active form of human Origin Recognition Complex and its ATPase motor module.
Tocilj, A., On, K.F., Yuan, Z., Sun, J., Elkayam, E., Li, H., Stillman, B., Joshua-Tor, L.(2017) Elife 6
- PubMed: 28112645 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.20818
- Primary Citation Related Structures:
5UJ7, 5UJ8, 5UJM - PubMed Abstract:
Binding of the Origin Recognition Complex (ORC) to origins of replication marks the first step in the initiation of replication of the genome in all eukaryotic cells. Here, we report the structure of the active form of human ORC determined by X-ray crystallography and cryo-electron microscopy. The complex is composed of an ORC1/4/5 motor module lobe in an organization reminiscent of the DNA polymerase clamp loader complexes. A second lobe contains the ORC2/3 subunits. The complex is organized as a double-layered shallow corkscrew, with the AAA+ and AAA+-like domains forming one layer, and the winged-helix domains (WHDs) forming a top layer. CDC6 fits easily between ORC1 and ORC2, completing the ring and the DNA-binding channel, forming an additional ATP hydrolysis site. Analysis of the ATPase activity of the complex provides a basis for understanding ORC activity as well as molecular defects observed in Meier-Gorlin Syndrome mutations.
- W. M. Keck Structural Biology Laboratory, Cold Spring Harbor, New York, United States.
Organizational Affiliation: