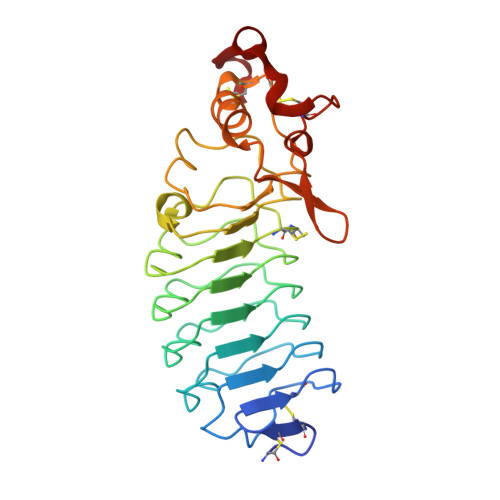
Structural Insights into VLR Fine Specificity for Blood Group Carbohydrates.
Collins, B.C., Gunn, R.J., McKitrick, T.R., Cummings, R.D., Cooper, M.D., Herrin, B.R., Wilson, I.A.(2017) Structure 25: 1667-1678.e4
- PubMed: 28988747 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2017.09.003
- Primary Citation Related Structures:
5UEI, 5UF1, 5UF4, 5UFB, 5UFC, 5UFD, 5UFF - PubMed Abstract:
High-quality reagents to study and detect glycans with high specificity for research and clinical applications are severely lacking. Here, we structurally and functionally characterize several variable lymphocyte receptor (VLR)-based antibodies from lampreys immunized with O erythrocytes that specifically recognize the blood group H-trisaccharide type II antigen. Glycan microarray analysis and biophysical data reveal that these VLRs exhibit greater specificity for H-trisaccharide compared with the plant lectin UEA-1, which is widely used in blood typing. Among these antibodies, O13 exhibits superior specificity for H-trisaccharide, the basis for which is revealed by comparative analysis of high-resolution VLR:glycan crystal structures. Using a structure-guided approach, we designed an O13 mutant with further enhanced specificity for H-trisaccharide. These insights into glycan recognition by VLRs suggest that lampreys can produce highly specific glycan antibodies, and are a valuable resource for the production of next-generation glycan reagents for biological and biomedical research and as diagnostics and therapeutics.
- Department of Integrative Structural and Computational Biology and the Skaggs Institute for Chemical Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: