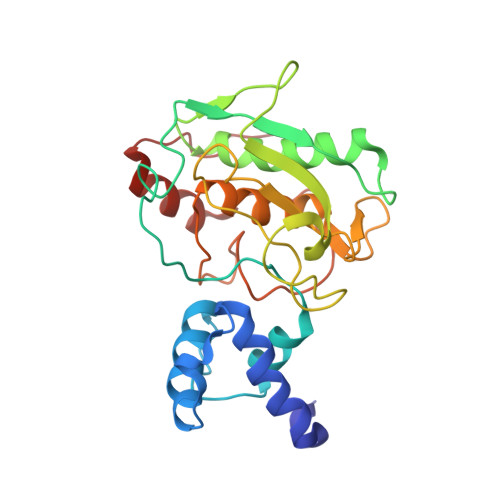
Glycan Activation of a Sheddase: Electrostatic Recognition between Heparin and proMMP-7.
Fulcher, Y.G., Prior, S.H., Masuko, S., Li, L., Pu, D., Zhang, F., Linhardt, R.J., Van Doren, S.R.(2017) Structure 25: 1100-1110.e5
- PubMed: 28648610 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2017.05.019
- Primary Citation Related Structures:
5UE2, 5UE5 - PubMed Abstract:
Heparan sulfate proteoglycans activate the matrix metalloproteinase-7 zymogen (proMMP-7) and recruit it in order to shed proteins from cell surfaces. This occurs in uterine and mammary epithelia, bacterial killing, lung healing, and tumor cell signaling. Basic tracks on proMMP-7 recognize polyanionic heparin, according to nuclear magnetic resonance and mutations disruptive of maturation. Contacts and proximity measurements guided docking of a heparin octasaccharide to proMMP-7. The reducing end fits into a basic pocket in the pro-domain while the chain continues toward the catalytic domain. Another oligosaccharide traverses a basic swath remote on the catalytic domain and inserts its reducing end into a slot formed with the basic C terminus. This latter association appears to support allosteric acceleration of proteolysis. The modes of binding account for extended, heterogeneous assemblies of proMMP-7 with heparinoids during maturation and for bridging to pro-α-defensins and proteoglycans. These associations support proteolytic release of activities at epithelial cell surfaces.
- Department of Biochemistry, University of Missouri, 117 Schweitzer Hall, Columbia, MO 65211, USA.
Organizational Affiliation: