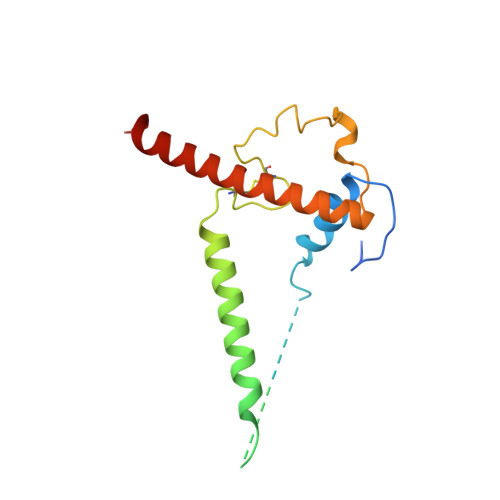
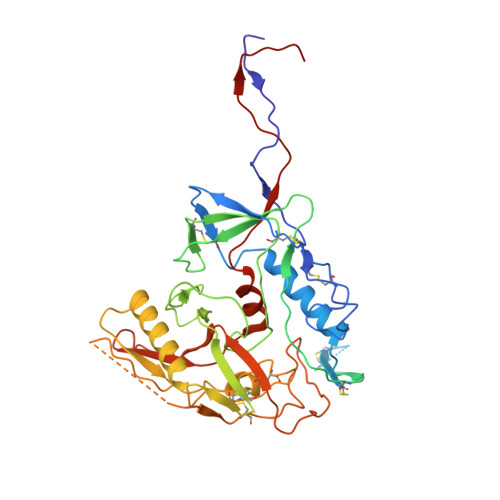
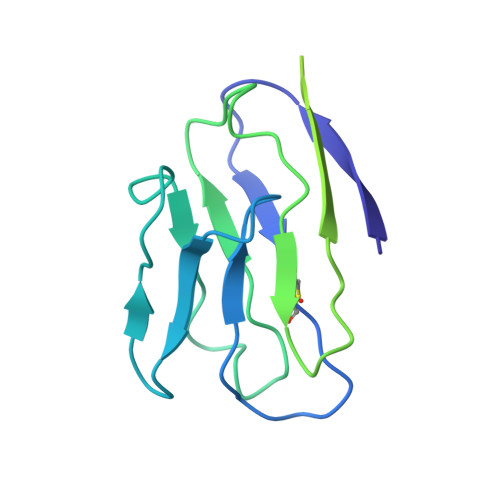
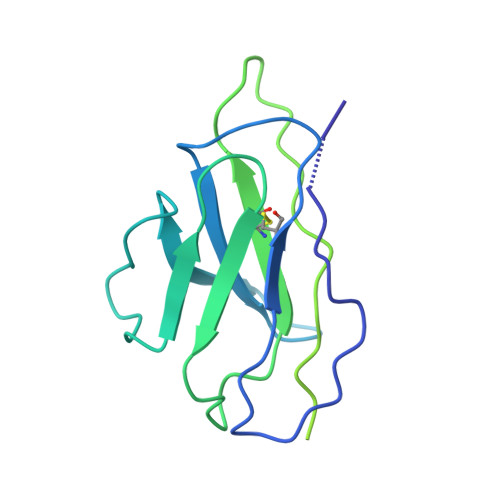
Cryo-EM structure of a CD4-bound open HIV-1 envelope trimer reveals structural rearrangements of the gp120 V1V2 loop.
Wang, H., Cohen, A.A., Galimidi, R.P., Gristick, H.B., Jensen, G.J., Bjorkman, P.J.(2016) Proc Natl Acad Sci U S A 113: E7151-E7158
- PubMed: 27799557 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1615939113
- Primary Citation Related Structures:
5THR - PubMed Abstract:
The HIV-1 envelope (Env) glycoprotein, a trimer of gp120-gp41 heterodimers, relies on conformational flexibility to function in fusing the viral and host membranes. Fusion is achieved after gp120 binds to CD4, the HIV-1 receptor, and a coreceptor, capturing an open conformational state in which the fusion machinery on gp41 gains access to the target cell membrane. In the well-characterized closed Env conformation, the gp120 V1V2 loops interact at the apex of the Env trimer. Less is known about the structure of the open CD4-bound state, in which the V1V2 loops must rearrange and separate to allow access to the coreceptor binding site. We identified two anti-HIV-1 antibodies, the coreceptor mimicking antibody 17b and the gp120-gp41 interface-spanning antibody 8ANC195, that can be added as Fabs to a soluble native-like Env trimer to stabilize it in a CD4-bound conformation. Here, we present an 8.9-Å cryo-electron microscopy structure of a BG505 Env-sCD4-17b-8ANC195 complex, which reveals large structural rearrangements in gp120, but small changes in gp41, compared with closed Env structures. The gp120 protomers are rotated and separated in the CD4-bound structure, and the three V1V2 loops are displaced by ∼40 Å from their positions at the trimer apex in closed Env to the sides of the trimer in positions adjacent to, and interacting with, the three bound CD4s. These results are relevant to understanding CD4-induced conformational changes leading to coreceptor binding and fusion, and HIV-1 Env conformational dynamics, and describe a target structure relevant to drug design and vaccine efforts.
- Division of Biology and Biological Engineering, California Institute of Technology, Pasadena, CA 91125.
Organizational Affiliation: