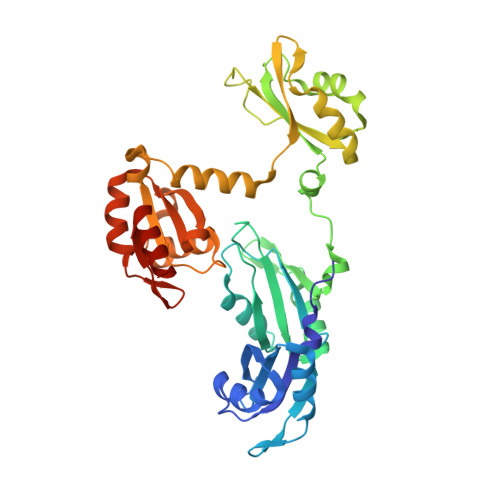
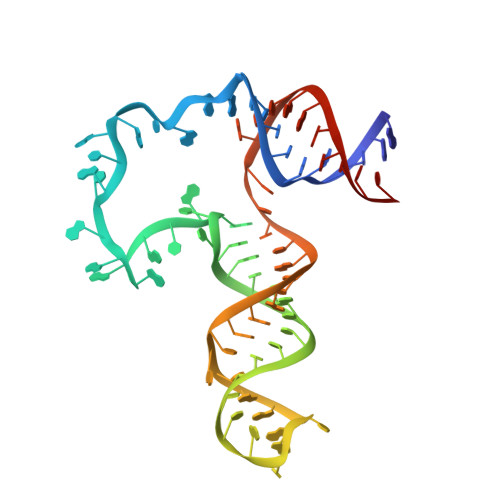
Structure and conformational plasticity of the U6 small nuclear ribonucleoprotein core.
Montemayor, E.J., Didychuk, A.L., Liao, H., Hu, P., Brow, D.A., Butcher, S.E.(2017) Acta Crystallogr D Struct Biol 73: 1-8
- PubMed: 28045380 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2059798316018222
- Primary Citation Related Structures:
5TF6 - PubMed Abstract:
U6 small nuclear RNA (snRNA) is a key component of the active site of the spliceosome, a large ribonucleoprotein complex that catalyzes the splicing of precursor messenger RNA. Prior to its incorporation into the spliceosome, U6 is bound by the protein Prp24, which facilitates unwinding of the U6 internal stem-loop (ISL) so that it can pair with U4 snRNA. A previously reported crystal structure of the `core' of the U6 small nuclear ribonucleoprotein (snRNP) contained an ISL-stabilized A62G mutant of U6 bound to all four RNA-recognition motif (RRM) domains of Prp24 [Montemayor et al. (2014), Nature Struct. Mol. Biol. 21, 544-551]. The structure revealed a novel topology containing interlocked rings of protein and RNA that was not predicted by prior biochemical and genetic data. Here, the crystal structure of the U6 snRNP core with a wild-type ISL is reported. This complex crystallized in a new space group, apparently owing in part to the presence of an intramolecular cross-link in RRM1 that was not observed in the previously reported U6-A62G structure. The structure exhibits the same protein-RNA interface and maintains the unique interlocked topology. However, the orientation of the wild-type ISL is altered relative to the A62G mutant structure, suggesting inherent structural dynamics that may facilitate its pairing with U4. Consistent with their similar architectures in the crystalline state, the wild-type and A62G variants of U6 exhibit similar Prp24-binding affinities and electrophoretic mobilities when analyzed by gel-shift assay.
- Department of Biochemistry, University of Wisconsin-Madison, Madison, WI 53706, USA.
Organizational Affiliation: