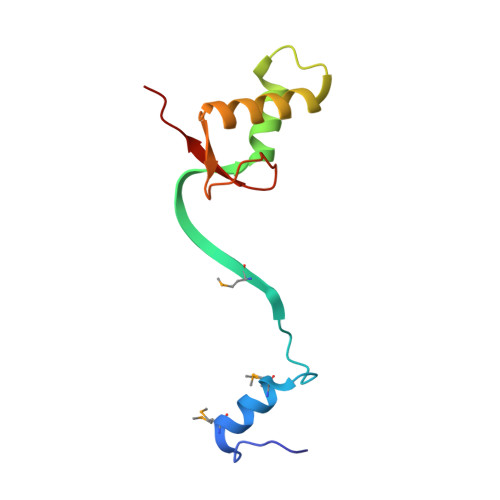
Wnt Signalosome Assembly by DEP Domain Swapping of Dishevelled.
Gammons, M.V., Renko, M., Johnson, C.M., Rutherford, T.J., Bienz, M.(2016) Mol Cell 64: 92-104
- PubMed: 27692984 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2016.08.026
- Primary Citation Related Structures:
5LNP, 5SUY, 5SUZ - PubMed Abstract:
Extracellular signals are often transduced by dynamic signaling complexes ("signalosomes") assembled by oligomerizing hub proteins following their recruitment to signal-activated transmembrane receptors. A paradigm is the Wnt signalosome, which is assembled by Dishevelled via reversible head-to-tail polymerization by its DIX domain. Its activity causes stabilization of β-catenin, a Wnt effector with pivotal roles in animal development and cancer. How Wnt triggers signalosome assembly is unknown. Here, we use structural analysis, as well as biophysical and cell-based assays, to show that the DEP domain of Dishevelled undergoes a conformational switch, from monomeric to swapped dimer, to trigger DIX-dependent polymerization and signaling to β-catenin. This occurs in two steps: binding of monomeric DEP to Frizzled followed by DEP domain swapping triggered by its high local concentration upon Wnt-induced recruitment into clathrin-coated pits. DEP domain swapping confers directional bias on signaling, and the dimerization provides cross-linking between Dishevelled polymers, illustrating a key principle underlying signalosome formation.
- MRC Laboratory of Molecular Biology, Cambridge Biomedical Campus, Francis Crick Avenue, Cambridge CB2 0QH, UK. Electronic address: melissag@mrc-lmb.cam.ac.uk.
Organizational Affiliation: