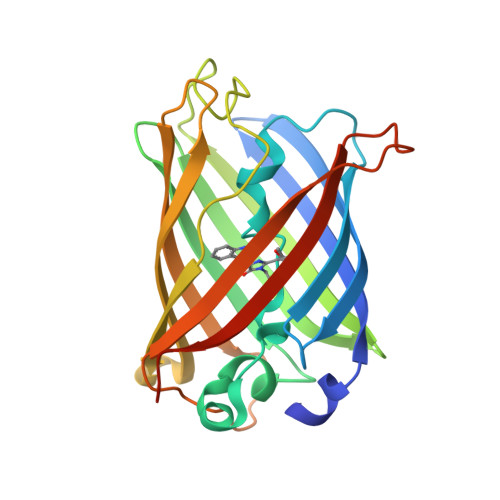
Chromophore Isomer Stabilization Is Critical to the Efficient Fluorescence of Cyan Fluorescent Proteins.
Gotthard, G., von Stetten, D., Clavel, D., Noirclerc-Savoye, M., Royant, A.(2017) Biochemistry 56: 6418-6422
- PubMed: 29148725 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.7b01088
- Primary Citation Related Structures:
5OX8, 5OX9, 5OXA, 5OXB, 5OXC - PubMed Abstract:
ECFP, the first usable cyan fluorescent protein (CFP), was obtained by adapting the tyrosine-based chromophore environment in green fluorescent protein to that of a tryptophan-based one. This first-generation CFP was superseded by the popular Cerulean, CyPet, and SCFP3A that were engineered by rational and random mutagenesis, yet the latter CFPs still exhibit suboptimal properties of pH sensitivity and reversible photobleaching behavior. These flaws were serendipitously corrected in the third-generation CFP mTurquoise and its successors without an obvious rationale. We show here that the evolution process had unexpectedly remodeled the chromophore environment in second-generation CFPs so they would accommodate a different isomer, whose formation is favored by acidic pH or light irradiation and which emits fluorescence much less efficiently. Our results illustrate how fluorescent protein engineering based solely on fluorescence efficiency optimization may affect other photophysical or physicochemical parameters and provide novel insights into the rational evolution of fluorescent proteins with a tryptophan-based chromophore.
- European Synchrotron Radiation Facility , F-38043 Grenoble, France.
Organizational Affiliation: