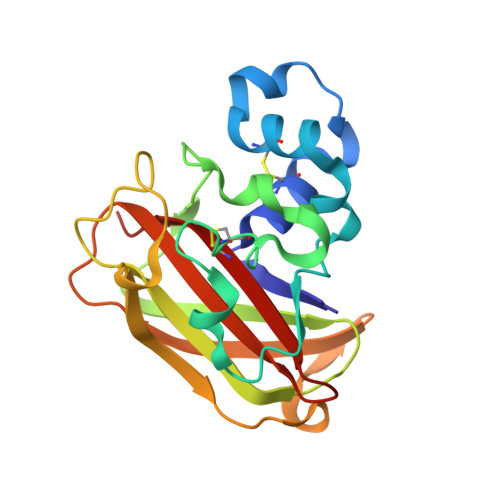
Structural determinants of bacterial lytic polysaccharide monooxygenase functionality.
Forsberg, Z., Bissaro, B., Gullesen, J., Dalhus, B., Vaaje-Kolstad, G., Eijsink, V.G.H.(2018) J Biological Chem 293: 1397-1412
- PubMed: 29222333 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M117.817130
- Primary Citation Related Structures:
5OPF - PubMed Abstract:
Bacterial lytic polysaccharide monooxygenases (LPMO10s) use redox chemistry to cleave glycosidic bonds in the two foremost recalcitrant polysaccharides found in nature, namely cellulose and chitin. Analysis of correlated mutations revealed that the substrate-binding and copper-containing surface of LPMO10s composes a network of co-evolved residues and interactions, whose roles in LPMO functionality are unclear. Here, we mutated a subset of these correlated residues in a newly characterized C1/C4-oxidizing LPMO10 from Micromonospora aurantiaca ( Ma LPMO10B) to the corresponding residues in strictly C1-oxidizing LPMO10s. We found that surface properties near the catalytic copper, i.e. side chains likely to be involved in substrate positioning, are major determinants of the C1:C4 ratio. Several Ma LPMO10B mutants almost completely lost C4-oxidizing activity while maintaining C1-oxidizing activity. These mutants also lost chitin-oxidizing activity, which is typically observed for C1/C4-oxidizing, but not for C1-oxidizing, cellulose-active LPMO10s. Selective loss in C1-oxidizing activity was not observed. Additional mutational experiments disclosed that neither truncation of the Ma LPMO10B family 2 carbohydrate-binding module nor mutations altering access to the solvent-exposed axial copper coordination site significantly change the C1:C4 ratio. Importantly, several of the mutations that altered interactions with the substrate exhibited reduced stability. This effect could be explained by productive substrate binding that protects LPMOs from oxidative self-inactivation. We discuss these stability issues in view of recent findings on LPMO catalysis, such as the involvement of H 2 O 2 Our results show that residues on the substrate-binding surface of LPMOs have co-evolved to optimize several of the interconnected properties: substrate binding and specificity, oxidative regioselectivity, catalytic efficiency, and stability.
- From the Faculty of Chemistry, Biotechnology, and Food Science, Norwegian University of Life Sciences (NMBU), 1432 Ås, Norway, zarah.forsberg@nmbu.no.
Organizational Affiliation: