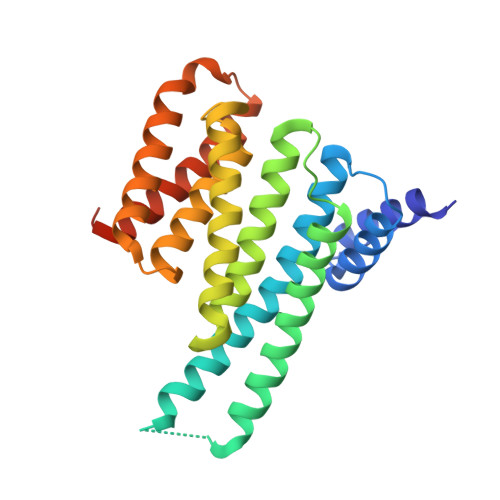
Chimeric 14-3-3 proteins for unraveling interactions with intrinsically disordered partners.
Sluchanko, N.N., Tugaeva, K.V., Greive, S.J., Antson, A.A.(2017) Sci Rep 7: 12014-12014
- PubMed: 28931924 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-017-12214-9
- Primary Citation Related Structures:
5OK9, 5OKF, 5OM0, 5OMA - PubMed Abstract:
In eukaryotes, several "hub" proteins integrate signals from different interacting partners that bind through intrinsically disordered regions. The 14-3-3 protein hub, which plays wide-ranging roles in cellular processes, has been linked to numerous human disorders and is a promising target for therapeutic intervention. Partner proteins usually bind via insertion of a phosphopeptide into an amphipathic groove of 14-3-3. Structural plasticity in the groove generates promiscuity allowing accommodation of hundreds of different partners. So far, accurate structural information has been derived for only a few 14-3-3 complexes with phosphopeptide-containing proteins and a variety of complexes with short synthetic peptides. To further advance structural studies, here we propose a novel approach based on fusing 14-3-3 proteins with the target partner peptide sequences. Such chimeric proteins are easy to design, express, purify and crystallize. Peptide attachment to the C terminus of 14-3-3 via an optimal linker allows its phosphorylation by protein kinase A during bacterial co-expression and subsequent binding at the amphipathic groove. Crystal structures of 14-3-3 chimeras with three different peptides provide detailed structural information on peptide-14-3-3 interactions. This simple but powerful approach, employing chimeric proteins, can reinvigorate studies of 14-3-3/phosphoprotein assemblies, including those with challenging low-affinity partners, and may facilitate the design of novel biosensors.
- A.N. Bach Institute of Biochemistry, Federal Research Center "Fundamentals of Biotechnology" of the Russian Academy of Sciences, 119071, Moscow, Russian Federation. nikolai.sluchanko@mail.ru.
Organizational Affiliation: