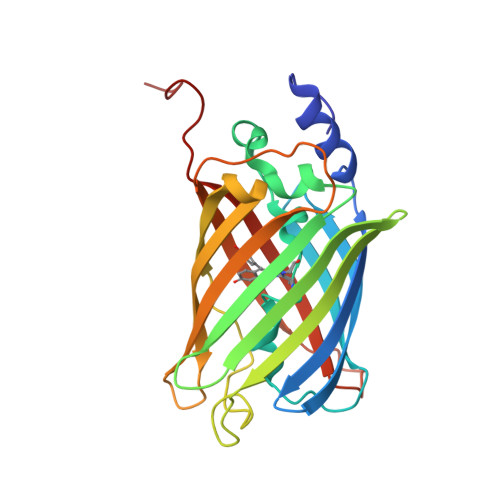
Chromophore twisting in the excited state of a photoswitchable fluorescent protein captured by time-resolved serial femtosecond crystallography.
Coquelle, N., Sliwa, M., Woodhouse, J., Schiro, G., Adam, V., Aquila, A., Barends, T.R.M., Boutet, S., Byrdin, M., Carbajo, S., De la Mora, E., Doak, R.B., Feliks, M., Fieschi, F., Foucar, L., Guillon, V., Hilpert, M., Hunter, M.S., Jakobs, S., Koglin, J.E., Kovacsova, G., Lane, T.J., Levy, B., Liang, M., Nass, K., Ridard, J., Robinson, J.S., Roome, C.M., Ruckebusch, C., Seaberg, M., Thepaut, M., Cammarata, M., Demachy, I., Field, M., Shoeman, R.L., Bourgeois, D., Colletier, J.P., Schlichting, I., Weik, M.(2018) Nat Chem 10: 31-37
- PubMed: 29256511 Search on PubMed
- DOI: https://doi.org/10.1038/nchem.2853
- Primary Citation Related Structures:
5O89, 5O8A, 5O8B, 5O8C - PubMed Abstract:
Chromophores absorb light in photosensitive proteins and thereby initiate fundamental biological processes such as photosynthesis, vision and biofluorescence. An important goal in their understanding is the provision of detailed structural descriptions of the ultrafast photochemical events that they undergo, in particular of the excited states that connect chemistry to biological function. Here we report on the structures of two excited states in the reversibly photoswitchable fluorescent protein rsEGFP2. We populated the states through femtosecond illumination of rsEGFP2 in its non-fluorescent off state and observed their build-up (within less than one picosecond) and decay (on the several picosecond timescale). Using an X-ray free-electron laser, we performed picosecond time-resolved crystallography and show that the hydroxybenzylidene imidazolinone chromophore in one of the excited states assumes a near-canonical twisted configuration halfway between the trans and cis isomers. This is in line with excited-state quantum mechanics/molecular mechanics and classical molecular dynamics simulations. Our new understanding of the structure around the twisted chromophore enabled the design of a mutant that displays a twofold increase in its off-to-on photoswitching quantum yield.
- Institut de Biologie Structurale, Univ. Grenoble Alpes, CEA, CNRS, F-38044 Grenoble, France.
Organizational Affiliation: