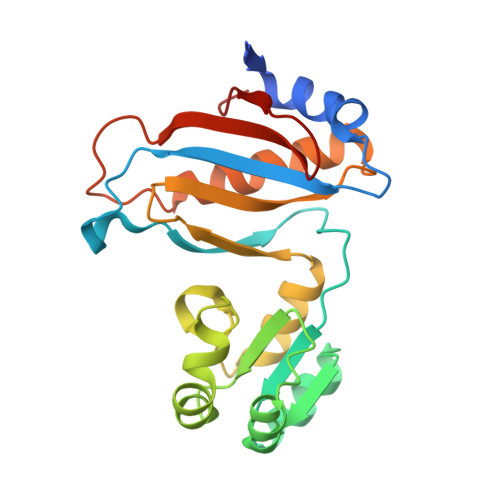
[Influence of Nonconserved Regions of L1 Protuberance of Thermus thermophilus Ribosome on the Affinity of L1 Protein to 23s rRNA].
Kostareva, O.S., Nevskaya, N.A., Tishchenko, S.V., Gabdulkhakov, A.G., Garber, M.B., Nikonov, S.V.(2018) Mol Biol (Mosk) 52: 106-111
- PubMed: 29512642 Search on PubMed
- DOI: https://doi.org/10.7868/S0026898418010147
- Primary Citation Related Structures:
5NPM - PubMed Abstract:
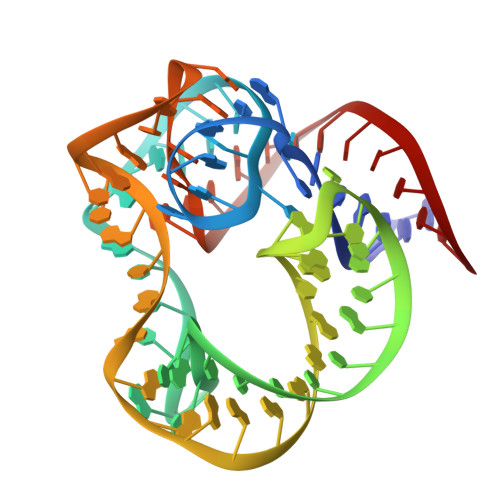
The L1 protuberance of the ribosome includes two domain ribosomal protein L1 and three helices of 23S rRNA (H76, H77, and H78) with interconnecting loops A and B. Helix 78 consists of two parts, i.e., H78a and H78b. A comparison of the available structural data of L1-RNA complexes with the obtained kinetic data made it possible to determine the influence of the nonconserved regions of Thermus thermophilus L1-protuberance on the mutual affinity of the L1 protein and 23S rRNA. It has been shown that the N-terminal helix of the protein and 78b helix of 23S rRNA are essential for the formation of an additional intermolecular contact, which is separated in the protein from the main site of L1-rRNA interaction by a flexible connection. This results in a rise in the TthL1-rRNA affinity. At the same time, the elongation of the 76 helix has no effect on rRNA-protein binding.
- Institute of Protein Research, Russian Academy of Sciences, Pushchino, Moscow oblast, 142290 Russia.
Organizational Affiliation: