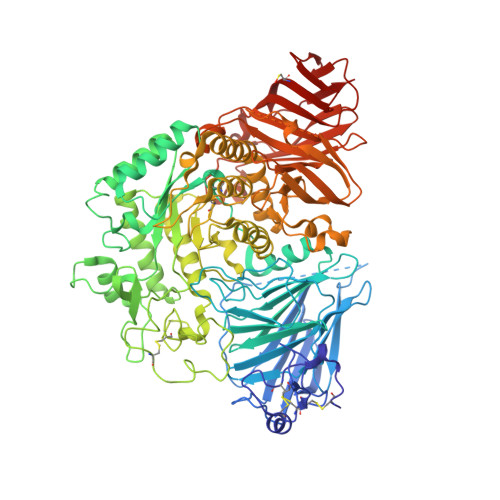
Structure of human lysosomal acid alpha-glucosidase-a guide for the treatment of Pompe disease.
Roig-Zamboni, V., Cobucci-Ponzano, B., Iacono, R., Ferrara, M.C., Germany, S., Bourne, Y., Parenti, G., Moracci, M., Sulzenbacher, G.(2017) Nat Commun 8: 1111-1111
- PubMed: 29061980 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-017-01263-3
- Primary Citation Related Structures:
5NN3, 5NN4, 5NN5, 5NN6, 5NN8 - PubMed Abstract:
Pompe disease, a rare lysosomal storage disease caused by deficiency of the lysosomal acid α-glucosidase (GAA), is characterized by glycogen accumulation, triggering severe secondary cellular damage and resulting in progressive motor handicap and premature death. Numerous disease-causing mutations in the gaa gene have been reported, but the structural effects of the pathological variants were unknown. Here we present the high-resolution crystal structures of recombinant human GAA (rhGAA), the standard care of Pompe disease. These structures portray the unbound form of rhGAA and complexes thereof with active site-directed inhibitors, providing insight into substrate recognition and the molecular framework for the rationalization of the deleterious effects of disease-causing mutations. Furthermore, we report the structure of rhGAA in complex with the allosteric pharmacological chaperone N-acetylcysteine, which reveals the stabilizing function of this chaperone at the structural level.
- Centre National de la Recherche Scientifique (CNRS), Aix-Marseille Univ, AFMB, 163 Avenue de Luminy, 13288, Marseille, France.
Organizational Affiliation: