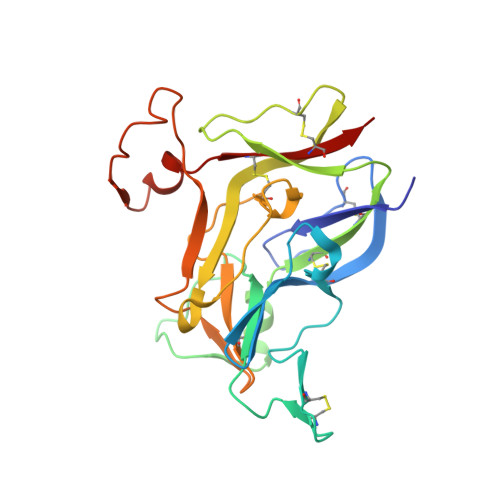
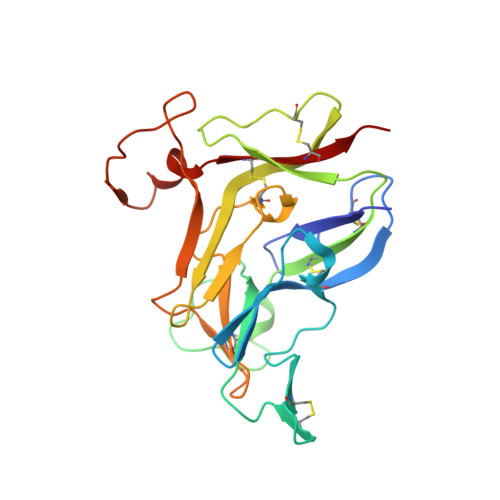
Structures of collagen IV globular domains: insight into associated pathologies, folding and network assembly.
Casino, P., Gozalbo-Rovira, R., Rodriguez-Diaz, J., Banerjee, S., Boutaud, A., Rubio, V., Hudson, B.G., Saus, J., Cervera, J., Marina, A.(2018) IUCrJ 5: 765-779
- PubMed: 30443360 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2052252518012459
- Primary Citation Related Structures:
5NAX, 5NAY, 5NAZ, 5NB0, 5NB1, 5NB2 - PubMed Abstract:
Basement membranes are extracellular structures of epithelia and endothelia that have collagen IV scaffolds of triple α-chain helical protomers that associate end-to-end, forming networks. The molecular mechanisms by which the noncollagenous C-terminal domains of α-chains direct the selection and assembly of the α1α2α1 and α3α4α5 hetero-oligomers found in vivo remain obscure. Autoantibodies against the noncollagenous domains of the α3α4α5 hexamer or mutations therein cause Goodpasture's or Alport's syndromes, respectively. To gain further insight into oligomer-assembly mechanisms as well as into Goodpasture's and Alport's syndromes, crystal structures of non-collagenous domains produced by recombinant methods were determined. The spontaneous formation of canonical homohexamers (dimers of trimers) of these domains of the α1, α3 and α5 chains was shown and the components of the Goodpasture's disease epitopes were viewed. Crystal structures of the α2 and α4 non-collagenous domains generated by recombinant methods were also determined. These domains spontaneously form homo-oligomers that deviate from the canonical architectures since they have a higher number of subunits (dimers of tetramers and of hexamers, respectively). Six flexible structural motifs largely explain the architectural variations. These findings provide insight into noncollagenous domain folding, while supporting the in vivo operation of extrinsic mechanisms for restricting the self-assembly of noncollagenous domains. Intriguingly, Alport's syndrome missense mutations concentrate within the core that nucleates the folding of the noncollagenous domain, suggesting that this syndrome, when owing to missense changes, is a folding disorder that is potentially amenable to pharmacochaperone therapy.
- Department of Biochemistry and Molecular Biology/ERI BIOTECMED, Universitat de València, Dr Moliner 50, Burjassot, 46100 Valencia, Spain.
Organizational Affiliation: