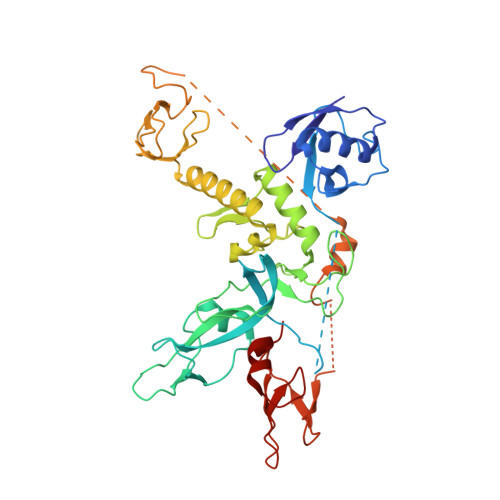
Parkin-phosphoubiquitin complex reveals cryptic ubiquitin-binding site required for RBR ligase activity.
Kumar, A., Chaugule, V.K., Condos, T.E.C., Barber, K.R., Johnson, C., Toth, R., Sundaramoorthy, R., Knebel, A., Shaw, G.S., Walden, H.(2017) Nat Struct Mol Biol 24: 475-483
- PubMed: 28414322 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.3400
- Primary Citation Related Structures:
5N2W, 5N38 - PubMed Abstract:
RING-between-RING (RBR) E3 ligases are a class of ubiquitin ligases distinct from RING or HECT E3 ligases. An important RBR ligase is Parkin, mutations in which lead to early-onset hereditary Parkinsonism. Parkin and other RBR ligases share a catalytic RBR module but are usually autoinhibited and activated via distinct mechanisms. Recent insights into Parkin regulation predict large, unknown conformational changes during Parkin activation. However, current data on active RBR ligases reflect the absence of regulatory domains. Therefore, it remains unclear how individual RBR ligases are activated, and whether they share a common mechanism. We now report the crystal structure of a human Parkin-phosphoubiquitin complex, which shows that phosphoubiquitin binding induces movement in the 'in-between RING' (IBR) domain to reveal a cryptic ubiquitin-binding site. Mutation of this site negatively affects Parkin's activity. Furthermore, ubiquitin binding promotes cooperation between Parkin molecules, which suggests a role for interdomain association in the RBR ligase mechanism.
- MRC Protein Phosphorylation and Ubiquitylation Unit, College of Life Sciences, University of Dundee, Dundee, UK.
Organizational Affiliation: