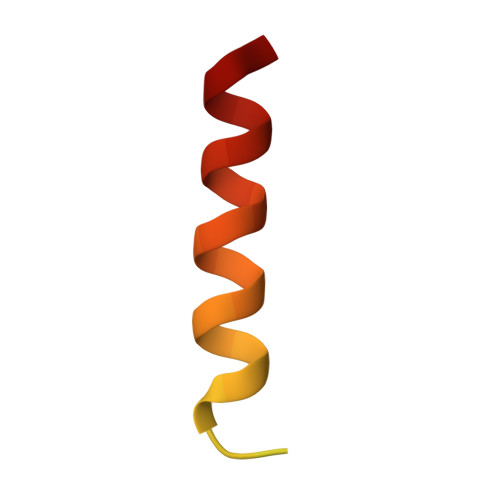
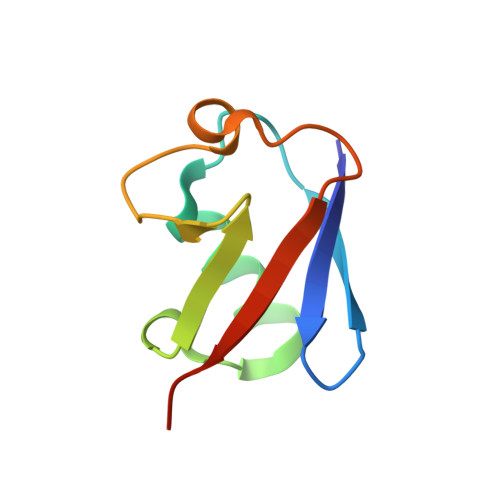
A single MIU motif of MINDY-1 recognizes K48-linked polyubiquitin chains.
Kristariyanto, Y.A., Abdul Rehman, S.A., Weidlich, S., Knebel, A., Kulathu, Y.(2017) EMBO Rep 18: 392-402
- PubMed: 28082312 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embr.201643205
- Primary Citation Related Structures:
5MN9 - PubMed Abstract:
The eight different types of ubiquitin (Ub) chains that can be formed play important roles in diverse cellular processes. Linkage-selective recognition of Ub chains by Ub-binding domain (UBD)-containing proteins is central to coupling different Ub signals to specific cellular responses. The motif interacting with ubiquitin (MIU) is a small UBD that has been characterized for its binding to monoUb. The recently discovered deubiquitinase MINDY-1/FAM63A contains a tandem MIU repeat (tMIU) that is highly selective at binding to K48-linked polyUb. We here identify that this linkage-selective binding is mediated by a single MIU motif (MIU2) in MINDY-1. The crystal structure of MIU2 in complex with K48-linked polyubiquitin chains reveals that MIU2 on its own binds to all three Ub moieties in an open conformation that can only be accommodated by K48-linked triUb. The weak Ub binder MIU1 increases overall affinity of the tMIU for polyUb chains without affecting its linkage selectivity. Our analyses reveal new concepts for linkage selectivity and polyUb recognition by UBDs.
- MRC Protein Phosphorylation and Ubiquitylation Unit, School of Life Sciences, University of Dundee, Dundee, UK.
Organizational Affiliation: